Researchers at Berkeley Lab and their collaborators who work on the Phenix software suite have published a new paper that summarizes how to determine three-dimensional macromolecular structures from three experimental methods: X-ray crystallography, neutron diffraction, and electron cryo-microscopy (cryo-EM). The article appeared in the journal Acta Crystallographica Section D: Structural Biology and is featured on the cover of the October 2019 issue.
MBIB Leadership Changes Announced
Molecular Biophysics and Integrated Bioimaging (MBIB) Division Director Paul Adams has announced a number of changes in the Division leadership, effective October 1. Earlier this summer, Corie Ralston agreed to serve as the Interim Director of the Molecular Foundry’s Biological Nanostructures Facility. She will step down as MBIB Division Deputy and remain as the Head of the Berkeley Center for Structural Biology (BCSB) at the Advanced Light Source (ALS).
BCSB Helps Elucidate Mechanism of Innate Immune Response
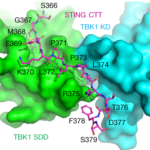
The crystallographic study of STING (stimulator of interferon genes), a transmembrane protein that plays a key role in innate immunity, in complex with TBK1 (serine/threonine-protein kinase), an enzyme that regulates the inflammatory response to foreign DNA, is extremely challenging due to weakly diffracting crystals. But thanks to the expertise of Berkeley Center for Structural Biology (BCSB) scientists, researchers from Texas A&M University (TAMU) were able to pinpoint the conserved motif of STING that mediates the recruitment and activation of TBK1. They published their results in Nature.
First Look at New Light Absorbing Protein
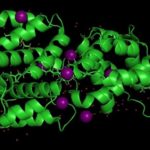
The Helical Carotenoid Protein 2 (HCP2) protein is an ancestor of proteins that are known to protect against damage caused by excess light exposure. Researchers in the laboratory of Cheryl Kerfeld, guest faculty in the Environmental Genomics & Systems Biology (EGSB) Division, are the first to structurally and biophysically analyze a protein from the HCP family. This HCP protein family was discovered recently by Kerfeld and the members of her lab, who are based in EGSB and at Michigan State University (MSU). To solve the molecular structure of HCP2, X-ray diffraction was measured at beam line 5.0.2 in the Berkeley Center for Structural Biology of the Advanced Light Source (ALS). The structure was refined using Phenix, a software suite for automated determination of molecular structures developed under the direction of Paul Adams, Molecular Biophysics and Integrated Bioimaging Division Director. Read more in the MSU-DOE Plant Research Laboratory news story.
Revealing the Shapes of Molecular Machines
Within each cell of the human body, thousands of molecular machines are at work. They transport nutrients and biochemicals into and out of our cells, build other tiny machines, and even move our cells around. To understand how these molecular machines work, scientists create three-dimensional pictures using electron cryomicroscopy (cryo-EM), catching these machines in different shapes that give insight into their function. Now researchers at Berkeley Lab and their international collaborators who write and distribute the Phenix software suite have developed a new set of computational tools for automated structure determination from cryo-EM data.
- « Previous Page
- 1
- …
- 3
- 4
- 5
- 6
- 7
- …
- 9
- Next Page »
Was this page useful?