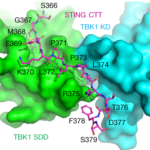
The crystallographic study of STING (stimulator of interferon genes), a transmembrane protein that plays a key role in innate immunity, in complex with TBK1 (serine/threonine-protein kinase), an enzyme that regulates the inflammatory response to foreign DNA, is extremely challenging due to weakly diffracting crystals. But thanks to the expertise of Berkeley Center for Structural Biology (BCSB) scientists, researchers from Texas A&M University (TAMU) were able to pinpoint the conserved motif of STING that mediates the recruitment and activation of TBK1. They published their results in Nature.
Antibody Uses Mimicry to Block SARS Coronavirus
To better understand how coronavirus antibodies work, a team of researchers at the University of Washington studied spike protein structures in complex with neutralizing antibody fragments isolated from SARS and MERS survivors. To visualize how the spike structures interact with the antibody fragments, they used a cryo-electron microscopy (cryo-EM) for the spikes, which are resistant to crystallization, and protein crystallography for the fragments. The high-resolution x-ray crystallography was performed at the Advanced Light Source (ALS) Beamline 5.0.1, part of the Berkeley Center for Structural Biology (BCSB).
BCSB Helps Characterize New Arsenic-based Antibiotic
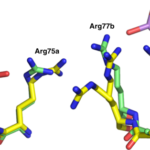
A newly-discovered arsenic-containing compound produced by a soil bacterium shows promise as a broad-spectrum antibiotic. In a paper published in the Nature journal Communications Biology, an international team of researchers demonstrated that arsinothricin (AST) is effective against many types of gram-negative and gram-positive bacteria. The effort was led by Barry Rosen of the Florida International University College of Medicine and Masafumi Yoshinaga of the National Agriculture and Food Research Organization (NARO) in Japan. Banumathi Sankaran, a research scientist in the Berkeley Center for Structural Biology (BSCB) at the Advanced Light Source (ALS), was an author on the paper.
First Look at New Light Absorbing Protein
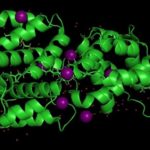
The Helical Carotenoid Protein 2 (HCP2) protein is an ancestor of proteins that are known to protect against damage caused by excess light exposure. Researchers in the laboratory of Cheryl Kerfeld, guest faculty in the Environmental Genomics & Systems Biology (EGSB) Division, are the first to structurally and biophysically analyze a protein from the HCP family. This HCP protein family was discovered recently by Kerfeld and the members of her lab, who are based in EGSB and at Michigan State University (MSU). To solve the molecular structure of HCP2, X-ray diffraction was measured at beam line 5.0.2 in the Berkeley Center for Structural Biology of the Advanced Light Source (ALS). The structure was refined using Phenix, a software suite for automated determination of molecular structures developed under the direction of Paul Adams, Molecular Biophysics and Integrated Bioimaging Division Director. Read more in the MSU-DOE Plant Research Laboratory news story.
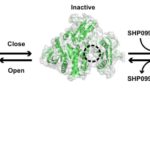
Locking Protein Structure to Close the Door on Cancer
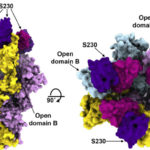
Mutations in the proteins that regulate cellular processes such as growth, division, and death are often linked to cancer and other diseases. The proper function of one of these proteins, SHP2, depends on maintaining equilibrium in a structural tug-of-war between an open (active) and a closed (inactive) arrangement. A team of researchers from Brandeis University performed X-ray crystallography at Advanced Light Source (ALS) Beamlines 8.2.1 and 8.2.2—part of the Berkeley Center for Structural Biology—to elucidate the structures of healthy and mutated forms of SHP2 and the dynamic interchange between their open and closed conformations, as well as how SHP2 interacts with certain cancer drugs.
Read more in this ALS Science Highlight.
- « Previous Page
- 1
- …
- 4
- 5
- 6
- 7
- 8
- …
- 12
- Next Page »
Was this page useful?