New crystallography finding by JBEI and GLBRC benefits bioenergy industry
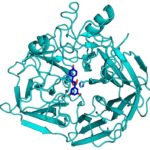 During the kraft process used to convert wood into wood pulp, the structural material lignin is partially converted into molecules like stilbene. Stilbenes are also naturally occurring in plants and some bacteria, and may play a role in plant pathogen resistance.
During the kraft process used to convert wood into wood pulp, the structural material lignin is partially converted into molecules like stilbene. Stilbenes are also naturally occurring in plants and some bacteria, and may play a role in plant pathogen resistance.
Currently, the deconstruction of plant biomass into cellulose and lignin is an expensive process. Lignin accounts for about 30 percent of plant cell wall carbon, and its conversion into chemicals or fuels could have a significant positive impact on the economics of processing lignocellulosic biomass. Enzymes capable of producing useful compounds from the breakdown of stilbenes and similar molecules could be employed for this. Collaborators from two of the Department of Energy Bioenergy Research Centers now have gained first-hand insight into how a stilbene cleaving oxygenase (SCO) carries out this unusual chemical reaction.