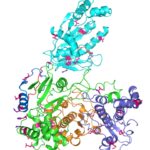
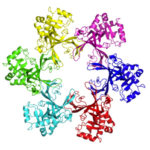
Zika virus is a mosquito-borne infectious disease linked to certain birth defects in infants in South and Central America and the United States. A Lawrence Berkeley National Laboratory (Berkeley Lab) researcher, Banumathi Sankaran, worked as part of a multi-institutional team led by Cheng Kao, professor at Indiana University, and Pingwei Li, associate professor at Texas A&M University (TAMU), to map a key viral protein called NS5. Necessary to virus reproduction, NS5 contains two enzyme activities: one reduces the body’s ability to mount an immune response against infection and the other helps start the genetic replication process.
Three Is Not a Crowd: Designed Metalloprotein Trimer Provides Stable Platform for Further Development
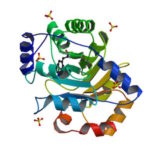 University of Washington (UW) researchers have designed a novel protein with properties that could lead to the generation of new photoactive proteins. This three-fold symmetric, self-assembling protein homotrimer contains a highly stable noncanonical amino acid. Noncanonical amino acids are not found among the 20 encoded amino acids in the body and can contain modifications to allow for new functionality. In this case, this amino acid contains a bipyridine group that chelates metal, thereby introducing new photochemical properties into the protein interface, and nucleating the formation of the homotrimer.
University of Washington (UW) researchers have designed a novel protein with properties that could lead to the generation of new photoactive proteins. This three-fold symmetric, self-assembling protein homotrimer contains a highly stable noncanonical amino acid. Noncanonical amino acids are not found among the 20 encoded amino acids in the body and can contain modifications to allow for new functionality. In this case, this amino acid contains a bipyridine group that chelates metal, thereby introducing new photochemical properties into the protein interface, and nucleating the formation of the homotrimer.
An article published last month in PNAS describes this work from a team of scientists led by David Baker at UW, which included Jose Henrique Pereira, Banumathi Sankaran, and Peter Zwart of the Molecular Biophysics & Integrated Bioimaging Division (MBIB). The MBIB scientists developed the crystal screen that was used to crystallize the novel protein and performed X-ray crystallography on Beamline 8.2.1 in the Berkeley Center for Structural Biology at the Advanced Light Source. Their X-ray crystallographic analysis of the homotrimer showed that the design process had near-atomic-level accuracy, demonstrating that computational protein design together with the utilization of noncanonical amino acids could be used to generate novel protein functions. These methods could be used to develop new therapeutics, biomaterials, and metalloproteins with useful optical or photochemical properties.
Biosciences Staff Honored with Director’s Awards
Several Biosciences Area personnel have been named as recipients of 2016 Berkeley Lab Director’s Awards. Yan Liang (Biological Systems & Engineering), Eva Nogales, and William Jagust (Molecular Biophysics & Integrated Bioimaging, MBIB) were honored with individual awards in Early Career, Scientific Achievement, and Societal Impact, respectively. Jill Fuss and Steven Yannone (MBIB) were the recipients of a team award in Technology Transfer for the launch of their company CinderBio. Jim Bristow (Biosciences Area Office, Trent Northen (Environmental Genomics & Systems Biology & Joint Genome Institute, JGI), and Susannah Tringe (JGI), along with Eoin Brodie and Peter Nico of the Earth and Environmental Sciences Area, were named in a team award in Service.
The Golden Time to Do Hybrid Methodologies
The work of Eva Nogales and Jennifer Doudna, Molecular Biophysics & Integrated Bioimaging Division faculty scientists, were highlighted in the August 18 Nature technology feature article, “Let the structural symphony begin,” a discussion of the current “golden time to do hybrid methodologies,” according to Nogales. Structural biologists, such as Nogales who was interviewed for this article, now have “the tools to tackle important questions about cells’ molecular machinery that would have been impossible to answer just a few years ago.” Combining these imaging methods – selecting from an impressive suite of different imaging techniques – is an increasingly popular approach in the field of structural biology, in which X-ray crystallography has been the premier method for more than a century. According to the article, while “the most powerful insights come from (these) hybrid methodologies that integrate images from several different tools, ” it also has its drawbacks. Read the full article here.
- « Previous Page
- 1
- …
- 3
- 4
- 5
Was this page useful?