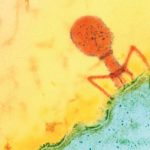
While sequencing gut bacteria from people in Bangladesh, Berkeley Lab’s Jillian Banfield discovered phages, viruses that infect and reproduce inside bacteria, twice as big as any previously found in humans. She and her colleagues found the snippets of megaphage DNA in a CRISPR segment of one type of bacteria, Prevotella, that is uncommon in people eating a high-fat, low-fiber Westernized diet. Banfield and her team named the clade of megaphages “Lak phage” after the Laksam Upazila area of Bangladesh where they were found.
Moving Beyond the Lab Bench
When researchers report developing a more efficient solar cell, or a technique that improves drug delivery, one of the inevitable follow-up questions they face is, “When will this be available to consumers?” In recent years, attempts to bridge the distance from lab bench to market have been promoted for researchers committed to seeing their work be applied to real world situations at universities, national laboratories and other institutions.
One such program is Fed Tech, launched in 2013 as part of the National Science Foundation’s Innovation Corps (I-Corps) program. Trent Northen and Peter Andeer, scientists in the Biosciences Area’s Environmental Genomics & Systems Biology (EGSB) Division at Lawrence Berkeley National Laboratory (Berkeley Lab), were part of the Fall 2018 cohort of Fed Tech Start-Up Studio, which culminated with a Pitch Day in late November. With entrepreneurs Rick Kjellberg and Jayan Rammohan, they were among 20 teams that participated in the eight-week program.
Updating the Genome Data Sharing Agreement
Nearly 20 years ago in Fort Lauderdale, Fla., genome data producers and data users came to an accord on the use of genome sequencing data released to the public domain. In particular, they agreed that the data was freely available for use and access by the scientific community before those data are used for publication. The Fort Lauderdale Agreement did not include defined policies on data usage, and has led to years of debate, such as whether or not there was a tacit acknowledgement that data generators would have the right of first publication on the data they produced and freely shared.
In a policy paper published January 25, 2019 in Science, 50 coauthors, with 54 unique affiliations from 18 countries, call for a “clear policy that protects public data from restrictions.” The international consortium includes Nikos Kyrpides of the Biosciences Area’s Environmental Genomics & Systems Biology (EGSB) Division at Lawrence Berkeley National Laboratory (Berkeley Lab).
Dub-seq Flagship Paper Published in Nature Communications
Dual barcoded shotgun expression library sequencing, or Dub-seq, is a novel high-throughput method for discovering gene function in microbes under various environmental conditions. It was developed by scientists in the Environmental Genomics and Systems Biology (EGSB) Division under the aegis of the Ecosystems and Networks Integrated with Genes and Molecular Assemblies (ENIGMA) program. In a seminal paper published January 18 in Nature Communications, the Dub-seq team presented details of the technique and proof-of-concept work showing that the approach is reproducible, economical, scalable, and identifies both known and novel gene functions.
New Ways to Control Bacterial Factories for Future Biotech Uses
A team of researchers led by Cheryl Kerfeld, faculty affiliate in the Environmental Genomics & Systems Biology (EGSB) Division, have developed a new method to manipulate miniature factories found in bacteria that could someday lead to new medical, industrial, or energy applications. The factories, called bacterial microcompartments – or BMCs – are found in bacteria all over the world.
- « Previous Page
- 1
- …
- 23
- 24
- 25
- 26
- 27
- …
- 45
- Next Page »
Was this page useful?