A critical component of the planned state-of-the-art macromolecular crystallography beamline Gemini has arrived at the Lab. The specially-designed in-vacuum undulator insertion device will yield higher photon flux compared to the Advanced Light Source’s superbend magnet and wiggler sources, bringing solutions to some of the most challenging projects in structural biology within reach.
Berkeley Lab Bioscientists Participate in CASP13
CASP is the Critical Assessment of Protein Structure Predictions, a biannual “competition” to determine which prediction algorithm generates the most accurate model. There are several categories in which models will be assessed, including accuracy, topology, and biological relevance. The SIBYLS beamline is participating to provide small-angle X-ray scattering (SAXS) data for––and judging for the first time––the “data-assisted” category. This CASP competition should lead to improvement in predicting protein-protein interfaces and complex structures.
ALS Passes the 7000-Protein Milestone
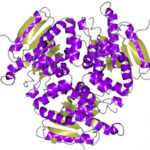
The Advanced Light Source hit a structural biology milestone in May 2018 with the help of their eight structural biology beamlines. Users of these beamlines have now collectively deposited over 7000 proteins into the Protein Data Bank (PDB), a worldwide, open-access repository of protein structures. The 7000th ALS protein structure (PBD accession number 6C7C) is an enzyme from Mycobacterium ulcerans (strain Agy99), solved with data from Beamline 5.0.2 in the Berkeley Center for Structural Biology. The enzyme is of interest to the researchers from the Seattle Structural Genomics Center for Infectious Disease (SSGID), whose mission is to obtain crystal structures of potential drug targets on the priority pathogen list of the National Institute of Allergy and Infectious Diseases (NIAID). Beamline 5.0.2, the first protein crystallography beamline at the ALS, came online in 1997. Read more in the ALS Feature.
ALS Enables Structural Determination of Respiratory Virus-Antibody Complex
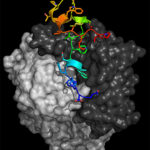
Using diffraction data collected at cryogenic temperature at the Advanced Light Source (ALS) Beamline 8.3.1, University of California Santa Cruz (UCSC) scientists, led by assistant professor of biomolecular engineering Rebecca DuBois, determined the three-dimensional atomic structure of a major surface protein of the respiratory syncytial virus (RSV): RSV G. Medical researchers have been trying to develop a vaccine for RSV, which causes serious respiratory disease in infants and older adults, for more than 50 years without success. But by leveraging the work of collaborators at Trellis Bioscience, who isolated protective human antibodies targeting RSV G, an attachment protein that allows the virus to stick to lung cells, the UCSC scientists were able to show that these protective antibodies target a section of the protein called the central conserved domain that is the same in all strains of the virus. The findings, published March 9 in Science Immunology, point to a promising route for designing a vaccine effective against a broad range of RSV strains. Read more from the UC Santa Cruz News Center.
Modified Antibody Clarifies Tumor-Killing Mechanisms
At the Advanced Light Source (ALS), researchers studied an antibody that was modified to activate a specific pathway of the immune system, demonstrating its value in killing tumor cells. The work provides a platform for disentangling the effects of different immune-system pathways and could lead to the design of improved cancer immunotherapies. The protein crystallography data in the study, which showed how an extreme twist in the modified antibody structure led to its selectivity, was collected at ALS Beamline 5.0.3 , which is part of the Biosciences’ Berkeley Center for Structural Biology; and at the Advanced Photon Source. Go to the ALS website to learn more.
- « Previous Page
- 1
- …
- 13
- 14
- 15
- 16
- 17
- …
- 19
- Next Page »
Was this page useful?