Trent R. Northen
Science Deputy, EGSB Division
Chemist Senior Scientist
Divisions
Environmental Genomics and Systems Biology
- Molecular EcoSystems Biology
Secondary Affiliation:
Biography
Metabolomics Program Lead
Deputy Division Director, Environmental Genomics and Systems Biology
Senior Scientist, Berkeley Lab
Education
B.S. in Chemical Engineering, University of California Santa Barbara;
Ph.D. in Chemistry and Biochemistry, Arizona State University;
Post-doc in Metabolomics, The Scripps Research Institute;
Executive Education, UC Berkeley Haas School of Business;
DOE Oppenheimer Science and Energy Leadership
Summary
Dr. Trent Northen is currently a Senior Staff Scientist at Berkeley Lab. Since joining Berkeley lab in 2008 Dr. Northen has had leadership roles within large DOE funded research programs using mass spectrometry for energy and environmental research including through the Joint BioEnergy Institute and the ENIGMA SFA project. In 2015, he founded what is now the JGI Metabolomics Program to provide advanced metabolomic methods available to JGI users. Dr. Northen’s research is focused on using metabolomics to link genomes with environments to understand how webs of microbes cycle carbon and sustain biomes via the development and application of advanced mass spectrometry approaches and model laboratory ecosystems (EcoFABs).
Recent Awards and Service
2019 R&D100 Award, 2016 Berkeley Lab Director Exceptional Achievement Award, 2014 DOE Early Career Award, 2013 R&D100 award, 2009 Presidential Award for Science and Engineering PECASE awarded by President Obama, A JBEI Excellence Award, multiple LBNL SPOT Awards, and two NIH SPORE Career Development Awards. Dr. Northen currently serves as the Deputy Director of the Environmental Genomics and Systems Biology at Lawrence Berkeley National Laboratory, he was the interim Division Director within this same organization during the formation of the Division in 2015-2016. Dr. Northen regularly chairs workshops, organizes conference sessions, and serves on review panels. At Berkeley Lab he serves on a number of committees and is the Strategy Mentor for the Berkeley Lab Biosciences Environmental Strategy.
Research Interests
Please see our lab’s website for more detail on our research.
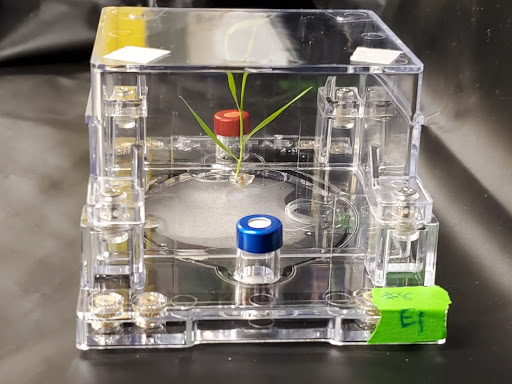
Summary: The combined impact of changing climate, population growth, environmental contamination, and soil degradation represent one of the greatest challenges facing humanity. Harnessing microbes will be a cornerstone of future sustainable agricultural and land management practices. Our laboratory uses mass spectrometry and fabricated ecosystem approaches to study the dynamic and reciprocal processes by which microbes interact to transform the organic molecules within their environment. Through the use of exometabolite profiling, we predict microbial metabolic webs in soils, sediment, and the rhizosphere for testing in fabricated ecosystems (EcoFABs) through the use of synthetic communities. Together these studies are providing insights into how microbes can be used to restore soil carbon, promote plant growth, and remove environmental contaminants.

Programs & Initiatives
Recent Publications
Related News
When Marine Algae Get Sick: How Viruses Shape Microbe Interactions
Researchers in the Environmental Genomics and Systems Biology Division collaborated on a study to better understand the role of viruses that infect photosynthetic phytoplankton in the marine food web.
Nurturing STEM Opportunities for Native Americans
A Berkeley Lab internship program aims to help increase Native American representation in graduate programs.
Introducing RhizoNet: AI-driven Plant Root Analysis
Berkeley Lab scientists from the Applied Mathematics and Computational Research (AMCR) and Environmental Genomics and Systems Biology (EGSB) Divisions developed RhizoNet, which harnesses the power of artificial intelligence (AI) to automate the process of root image analysis with exceptional accuracy.