Naomi S. Ginsberg
Chemist Faculty Scientist
Biography
Naomi S. Ginsberg is an Associate Professor of Chemistry and Physics at University of California, Berkeley and a Faculty Scientist in the Materials Sciences and Molecular Biophysics and Integrated Imaging Divisions at Lawrence Berkeley National Laboratory, where she has been since 2010. She currently focuses on elucidating the electronic and molecular dynamics in a wide variety of soft electronic and biological materials by devising new electron and optical imaging modalities that enable characterization of fast and ultrafast processes at the nanoscale and as a function of their heterogeneities. Naomi received a B.A.Sc. degree in Engineering Science from the University of Toronto in 2000 and a Ph.D. in Physics from Harvard University in 2007, after which she held a Glenn T. Seaborg Postdoctoral Fellowship at Lawrence Berkeley National Lab. Her background in chemistry, physics, and engineering has previously led her to observe initiating events of photosynthesis that take place in a millionth billionth of a second and to slow, stop, and store light pulses in some of the coldest atom clouds on Earth. She is the Berkeley lead of STROBE, a multi-university NSF Science and Technology Center devoted to imaging science, a member of the Kavli Energy Nanoscience Institute at Berkeley, and the recipient of a David and Lucile Packard Fellowship in Science and Engineering (2011), a DARPA Young Faculty Award (2012), an Alfred P. Sloan Foundation Fellowship (2015), and a Camille Dreyfus Teacher-Scholar Award (2016) in addition to a series of teaching awards in the physical sciences. In 2017-18 she was a Miller Professor for Basic Research in Science at UC Berkeley and was designated a Kavli Fellow.
Research Interests
We are pushing the limits of spatially resolved spectroscopy and time resolved microscopy in multiple modalities, tailored to answer fundamental and challenging questions that span chemistry, physics, and biology.
- How can we investigate the optical properties of soft matter and biological systems well below the diffraction limit? Can the nanoscale dynamics of both matter and energy in these systems be studied non-invasively in real-time?
- How does the local morphology of organic electronics affect their exciton dynamics? What is the relationship between local electron dynamics and overall device performance?
- How can we learn from the remarkable efficiency of photosynthesis to guide the design and optimization of biomimetic light harvesting systems? How can energy flow be manipulated?
A common theme in our work is to investigate light-matter interactions in the near- and far-field, on ultrafast time scales, with light and electron optics, in vacuum and in the condensed phase. Please consult the Research page for additional detail.
Recent Publications
Related News
Young Biosciences Researchers Rub Elbows with Nobel Company
MBIB graduate students Margaret Doyle and Christian Tanner were among 27 highly accomplished young UC scientists selected as fellows to the prestigious 2024 Lindau Nobel Nobel Laureate Meeting in Germany where they mingled with Nobel laureates.
Naomi Ginsberg Elected Fellow of the American Physical Society
Naomi Ginsberg, a faculty scientist in the Molecular Biophysics and Integrated Bioimaging (MBIB) Division, is among the 2021 class of Fellows elected by the American Physical Society (APS). The APS Fellowship Program recognizes members who have made exceptional contributions to the physics enterprise in research, applications, leadership, service, or education. Ginsberg, who is also a UC Berkeley associate professor of chemistry, was cited for her innovative development of spatiotemporally resolved imaging and spectroscopy methods—as well as their applications in elucidating energy transport in hierarchical and heterogeneous materials and in the formation and transformation of said materials.
Tracking Energy Flow in Light-harvesting Systems on Native Nanometer and Picosecond Scales
In the first trillionths of a second after sunlight hits a photosynthetic organism, the energy that is absorbed flows through a dense network of protein-bound chlorophyll molecules to a dedicated location where it is converted to electric charges. This is the first step in a series of events that ultimately drives the formation of sugar and starch to store energy in chemical bonds. “This migration is the triggering event that leads to all of the oxygen that we breathe, all of the food that we have, and we really don’t understand why this part of photosynthesis works as well as it does. For every photon of light that’s absorbed, you can expect some biochemical action to occur. That efficiency is really remarkable,” says Naomi Ginsberg, a faculty scientist in the Molecular Biophysics and Integrated Bioimaging (MBIB) Division who has a secondary affiliation in Materials Sciences and is also a UC Berkeley associate professor of Chemistry and Physics. Ginsberg and her colleagues devised a way to measure migration efficiency, and they describe the method in Nature Materials in November 2017.
Research Interests
My lab is currently working to improve two aspects of single-particle cryo-EM where there still remains a large gap between the current state-of-the-art and what is physically achievable. The first aspect is the way in which thin specimens are spread on EM grids, prior to rapid vitrification. We are developing a number of structure-friendly ways to immobilize particles onto affinity grids, in order to prevent their adsorption at the air-water interface. At the same time, we are developing new approaches to remove as much buffer as possible while avoiding that the air-water interface touches the now-immobilized particles. The second aspect is to use an intense, focused standing wave of light as a phase plate for the electron microscope, in order to provide in-focus image contrast that is much closer to the theoretical limit than what is currently possible.
Recent Publications
Related News
Congratulations to Biosciences Area Director’s Award Recipients
Numerous Biosciences Area personnel are among the 2021 Berkeley Lab Director’s Awards honorees. This annual program recognizes outstanding contributions by employees to all facets of Lab activities. A complete list of winners can be found here. The 10th annual Director’s Awards ceremony will take place on November 18 at noon.
Congratulations to Biosciences Area Director’s Award Recipients
Numerous Biosciences Area personnel are among the 2020 Berkeley Lab Director’s Awards honorees. This annual program recognizes outstanding contributions by employees to all facets of Lab activities. A complete list of winners can be found here. The ninth annual Director’s Awards ceremony will take place (virtually) on November 12 at 3 PM.
Glaeser Honored with Glenn T. Seaborg Medal
Robert Glaeser, senior scientist in the Molecular Biophysics & Integrated Bioimaging Division, was awarded the Glenn T. Seaborg Medal by the Department of Chemistry & Biochemistry at the University of California, Los Angeles (UCLA). At a symposium held on November 1o, Glaeser and Richard Henderson, Nobel Laureate in Chemistry 2017, were recognized for their crucial contributions to the science of electron cryo-microscopy.
Biography
Jay T. Groves received his B.S. degree in Physics and Chemistry from Tufts University, and then went on to complete his Ph.D. in Biophysics with Professors Steven Boxer and Harden McConnell at Stanford University. He then spent a year as a visiting scholar at Academia Sinica in Taipei, Taiwan before becoming the Division Director’s Fellow in the Physical Biosciences Division at Lawrence Berkeley National Laboratory. In 2001 he joined the Chemistry Department at UC Berkeley as an Assistant Professor. He was promoted to Associate Professor in 2007 and Professor in 2010. In 2008 Professor Groves was appointed as a Howard Hughes Medical Institute Investigator. He has received the Burroughs Wellcome Career Award in the Biomedical Sciences (2000), the Searle Scholars Award (2002), the MIT TR100 (2003), the Beckman Young Investigator Award (2004), and the NSF CAREER Award (2005). He has served as an Associate Editor of the Annual Reviews of Physical Chemistry since 2006.
Research Interests
Professor Groves is primarily interested in role of spatial organization in biochemical reaction systems. Living cells are not at all well-mixed reaction chambers. Rather, the molecular processes of life occur in elaborate spatial patterns. This interplay between spatial organization and the chemical reactions themselves in living systems adds a fascinating new dimension to chemistry that is rarely encountered outside of biology. Specific research in Professor Groves’ laboratory focuses on how spatial organization influences signal transduction processes at the cell membrane. The research methods combine techniques in optical microscopy and spectroscopy with materials fabrication methods and cell biology. This integrated approach enables the direct observation and physical manipulation of living reaction systems, down to the single molecule level. The conceptual approach is rooted in physics and physical chemistry, with the overarching goal of developing a quantitative and mechanistic understanding biochemical processes in living systems.
Recent Publications
Related News
How T Cells Tune Out Fake Signals: Phase Transition Timing Is Everything
A group of Berkeley Lab and UC Berkeley physical chemists led by Jay Groves, faculty scientist in Molecular Biophysics and Integrated Bioimaging (MBIB), has—for the first time—imaged the process by which an individual immune system molecule is switched on in response to a signal from the environment. This breakthrough led to the discovery that the immune system activation process involves hundreds of proteins suddenly coming together to form a linked network through a process known as phase transition. Critically, the process has a built in time delay which allows the cell to distinguish a genuine receptor stimulation from background chemical noise. The work is described in a paper recently published in the journal Science.

Building: Stanley Hall, Room 274D
Phone: (510) 495-2116
OHallatschek@lbl.gov
http://hallatscheklab.berkeley.edu/
Links
Research Interests
We are trying to understand how collective patterns of self-organization emerge from the joint actions of heterogeneous individuals. The phenomena studied include evolutionary adaptation, random genetic drift, epidemic spreading, collective motion, synchronization and jamming. Although these phenomena occur in many complex systems, our experimental efforts are focused primarily on microbial systems that we can study in our wet lab. Our key theoretical challenge is to identify essential dynamical building blocks and to predict how these conspire to generate the complex dynamical patterns observed at the population level. See publications for a detailed overview of our research.
Recent Publications
Divisions
Biological Systems and Engineering
- Process Engineering & Analytics
Secondary Affiliation:
Molecular Biophysics and Integrated Bioimaging
Biography
Amy E. Herr is the Lester John & Lynne Dewar Lloyd Distinguished Professor of Bioengineering at the University of California, Berkeley and a Chan Zuckerberg (CZ) Biohub Investigator. Prof. Herr joined UC Berkeley as Assistant Professor of Bioengineering in 2007, was promoted to Associate Professor with tenure in 2012, and promoted to Full Professor in 2015. Prior to joining UC Berkeley, she was a staff member in the Biosystems Research Group at Sandia National Laboratories (Livermore, CA; 2002-2007). She earned her PhD in Mechanical Engineering at Stanford with Profs. Tom Kenny & Juan Santiago as an NSF Graduate Research Fellow, an MS in Mechanical Engineering also from Stanford, and a BS in Engineering & Applied Science from Caltech.
Professor Herr is an elected Fellow of the National Academy of Inventors and the American Institute of Medical and Biological Engineering (AIMBE), a Board Member of the Chemical & Biological Microsystems Society (CBMS) which oversees the microTAS conferences, is a standing member of the NIH Nanotechnology Study Section, and is an Advisory Board Member for the UCSF Rosenman Institute and the journals Analytical Chemistry and ACS Sensors. She has served as a Co-Director of the Cold Spring Harbor Laboratory’s Single Cell Analysis summer course (2015 & 2016), both Chair (2009) and Vice-chair (2007) of the Gordon Research Conference (GRC) on the Physics & Chemistry of Microfluidics. She is faculty advisor to the UC Berkeley chapter of the Society of Women Engineers (SWE) and the Graduate Women in Engineering (GWE).
Professor Herr’s research has been recognized by: the 2016 Mid-career Achievement Award from the American Electrophoresis Society, the 2015 Georges Guiochon Faculty Fellow from HPLC, the 2012 Young Innovator Award from Analytical Chemistry/CBMS, the 2012 Ellen Weaver Award from the Association for Women in Science (AWIS, for mentoring), a 2011 NSF CAREER award, a 2010 NIH New Innovator Award, a 2010 Alfred P. Sloan Research Fellowship in chemistry, a 2010 New Investigator Award in Analytical Chemistry from Eli Lilly & Co., a 2009 Defense Advanced Research Projects Agency (DARPA) Young Faculty Award, a 2009 Hellman Family Faculty Fund Award from UC Berkeley, a 2008 Regents’ Junior Faculty Fellowship from the University of California. Professor Herr has also been recognized by the 2012 Outstanding Instructor Award in Bioengineering (Bioengineering Honor Society student vote) and a 2007 Outstanding Mentor Award from Sandia National Laboratories.
Research Interests
Scale-dependent Phenomena Underpinning Technology Development
Large-scale study of protein structure, function, and expression (proteomics) is instrumental to molecular biomarker discovery. Due to the constantly changing nature of protein expression and state, these profiles are notoriously difficult to study. High-resolution analytical assays such as two-dimensional electrophoresis and mass spectrometry have proven essential to proteomics; nevertheless, these information-rich methods can be slow and labor intensive. With these considerations in mind, our group is developing techniques, implemented via microfluidic technologies, as a means to achieve a rapid, yet still quantitative, assessment of protein expression & state variations in complex samples.
Biomarker Validation
In spite of significant advances in proteomic technology, few new protein biomarkers have emerged from the proteomic discovery pool, progressed though the scrutiny of validation studies, and become incorporated in diagnostic tools. The long-term goal of our work is development of flexible instruments for the rapid validation of putative disease-specific biomarkers in promising diagnostic fluids. An urgent need exists for robust bioanalytical capability that delivers high-throughput validation of putative biomarkers, thus allowing subsequent incorporation of validated markers into diagnostics. To achieve this aim, our group employs nascent microfluidic technologies to seamlessly integrate complex sample preparation, sample handling, and quantitative bioanalytical assays into tools amenable to automation.
Clinical & Point-of-Care Diagnostics
Appropriate, effective biomolecular analysis mechanisms are identified for diagnostic development based upon the physicochemical characteristics of putative, disease-specific biomarkers. Most disease states are complex — diagnosis & monitoring require more than simple binary detection of a small set of proteins. To compound the difficulty in assessing disease state, analytical grade quantitation and specificity are difficult to achieve as part of a disease diagnostic, especially diagnostics employed in near-patient environments. Consequently, our group is exploring the use of electrophoretic microfluidic formats, as such formats have been demonstrated to allow rapid, analytical-grade quantitation of small sample volumes through enhanced resolving power and high-efficiency operation.
Recent Publications
Related News
Chan Zuckerberg Biohub Network Names Herr CTO
The Chan Zuckerberg Biohub Network has tapped Amy Herr, faculty engineer in the Biological and Systems Engineering Division, to serve as Chief Technology Officer. She is charged with advancing technologies to observe, measure, and analyze human biology in action.
Congratulations 2021 Chan Zuckerberg Biohub Investigators
Four faculty scientists in the Biosciences Area were included in The Chan Zuckerberg Biohub Investigator Program, awarding $21 million to 21 University of California, Berkeley researchers.
Amy Herr to Head UC Berkeley’s Bakar BioEnginuity Hub
UC Berkeley has announced a new campus initiative, the Bakar BioEnginuity Hub (BBH), that aims to launch the world-changing startups of today, while cultivating the innovative leaders of tomorrow. Opening this fall, BBH will focus on people working at the convergence of the life sciences with the physical, engineering, and data sciences. Amy Herr, a faculty engineer in the Biological Systems and Engineering (BSE) Division and UC Berkeley professor of bioengineering, has been named executive director of BBH.
Research Interests
The Holmes Lab brings techniques from machine learning, statistical linguistics, phylogenetics, and web development to bear on the interpretation and analysis of genomic data. Examples include the application of context-free grammars to understanding DNA and RNA structure; the use of phylogenetic methods in genome annotation, and to detect recombination breakpoints; the development of machine learning algorithms for bioinformatics models; the reconstruction of insertion, deletion and transposition events in genome evolutionary histories; statistical algorithms for metagenomics species distribution analysis; and dynamic-HTML web applications for collaborative genomic data analysis.
Recent Publications

Building: Advanced Light Source Building, Room 2108
Phone: 510-486-4587
Fax: 510-486-5298
JMHolton@lbl.gov
Biography
Dr. Holton is currently an Associate Adjunct Professor at the University of California San Francisco with a Faculty Associate appointment at Lawrence Berkeley National Laboratory, where he serves as the Director for the Macromolecular Crystallography (MX) X-ray Diffraction Beamline 8.3.1 at the Advanced Light Source. He is an expert in optimizing sample preparation and diffraction techniques with particular focus on radiation damage, detector performance and x-ray scattering physics. He has written several absolute-scale simulators for both “conventional” MX and femtosecond nanocrystallography that have been instrumental in designing these experiments.
Dr. Holton earned a B.S., in Biology from the California Institute of Technology and his Ph.D., in Molecular and Cell Biology from the University of California at Berkeley.
Recent Publications
Related News
Congratulations to Biosciences Area Director’s Award Recipients
Several Biosciences Area personnel are among the 2024 recipients of Berkeley Lab Director’s Achievement Awards. The program recognizes outstanding contributions by employees to all aspects of Lab activities.
Biosciences Area FY24 LDRD Projects
The projects of 21 Biosciences Area scientists and engineers received funding through the FY24 Laboratory Directed Research and Development (LDRD) program.
Researchers Capture Elusive Missing Step in Photosynthesis
After decades of effort, scientists have revealed atomic-scale details of the water splitting step of photosynthesis, the chemical process that generates the air we breathe. The latest work adds to our understanding of photosynthesis and will aid the development of fully renewable alternative energy sources.

Building: Advanced Light Source Building , Room 2136
Phone: (510) 486-5378
Fax: 510-486-5298
GLHura@lbl.gov
Research Interests
Mechanisms of biological macromolecules inspire nanoscale engineering strategies and provide insights into disease.
The speed of genomic sequencing has rapidly increased; opening up 3 billion years of evolutionary engineering, new disease treatment opportunities and providing a perspective on the complex networks involved in most biological processes. Enabled by high throughput approaches, rather than focusing on a single system or pathway I study multiple pathways and processes. I work with the view that in cellular networks there are only a few degrees of separation between the actions of any two molecules. Of particular interest are hub proteins which, through multiple interactions or adopting specific conformations, signal alternate cellular outcomes. By developing an intuition in diverse bio-macromolecular systems I also work on engineering macromolecules for new functions.
Building intuition on the large networks and multi-level feedback loops in cellular systems requires many measurements which current capabilities cannot deliver.
An understanding of mechanism has fallen behind the rate at which new molecules of interest are being identified. I utilize and develop high throughput solution based techniques to characterize the conformations biomolecules adopt in the many contexts they encounter. A primary technique has been small angle X-ray scattering or SAXS. X-rays provide access to high resolution. New light sources provide exponentially increasing power. The two aspects combined provide insights into conformations of a macromolecule in high throughput. I also develop approaches to combine crystallographic results with SAXS. Crystallography is low throughput but provides un-paralleled resolution. SAXS provides access to conformational changes in high throughput.
Publications
Related News
SIBYLS Team Recognized with Innovative Instrumentation Award
The Structurally Integrated BiologY for the Life Sciences (SIBYLS) team received the 2025 Klaus Halbach Award for Innovative Instrumentation at the 2025 Advanced Light Source (ALS) User Meeting in August.
Time-Resolved SAXS Screen of Small-molecule Drug Candidates
A team of researchers developed a high-throughput drug-discovery workflow leveraging time-resolved small-angle X-ray scattering (SAXS) capabilities at the Advanced Light Source’s (ALS) Structurally Integrated Biology for the Life Sciences (SIBYLS) beamline to identify small molecules capable of activating biomolecular dynamics associated with a desired therapeutic outcome.
University of Duisburg-Essen Delegation Explores Collaborative Opportunities with Lawrence Berkeley National Laboratory
Building on a Memorandum of Understanding signed between UDE and Berkeley Lab researchers, a kick-off meeting focused on future collaborations in the fields of genomics, structural biology, bioimaging, and water research.

Building: Life Sciences Addition, Room 271
Phone: (510) 642-9853
EYIsacoff@lbl.gov
https://vcresearch.berkeley.edu/faculty/ehud-isacoff
Links
Research Interests
Our lab’s research is focused in four intersecting and complementary areas: mechanisms of ion channel function, synapse development and plasticity, neural circuit function, and the design of novel probes for the optical detection of neuronal signaling.
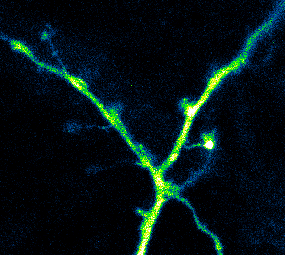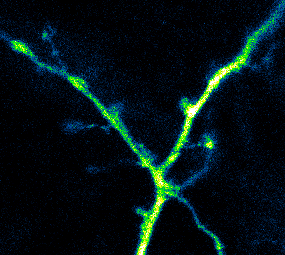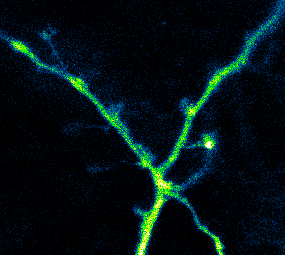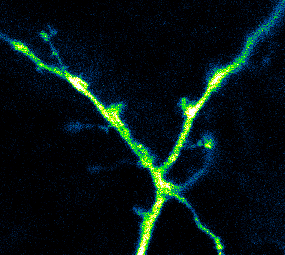
Developing Tools to Study Neurons
Elucidating structural and functional connectivity between neurons is one of today’s greatest challenges of systems neuroscience. We have developed a family of genetically-encoded photoactivatable calcium sensors that allow us to simultaneously visualize the morphology and activity of individual neurons, including fine structures such as dendritic spines.
Recent Publications
Related News
Biosciences Researchers Honored by the National Academy of Sciences
Three scientists affiliated with the Biosciences Area have been recognized by the National Academy of Sciences (NAS), one as the recipient of an NAS award and two as newly elected members. On Sunday, the NAS formally presented its 2018 NAS Award in Chemical Sciences to Jennifer Doudna, a faculty biochemist in the Molecular Biophysics and Integrated Bioimaging (MBIB) Division. Judith Campisi, a biochemist affiliated with Biological Systems and Engineering Division, Ehud “Udi” Isacoff, an MBIB faculty biologist, are among the group of 84 new members elected to the NAS.
DARPA Awards $21.6M to Develop Optogenetic ‘Read-Write’ Neural Interface
Ehud Isacoff of the Molecular Biophysics and Integrated Bioimaging (MBIB) Division is the project lead on a $21.6 million grant awarded to UC Berkeley as part of the Defense Advanced Research Projects Agency’s (DARPA’s) Neural Engineering System Design program. The team led by Isacoff, director of the Helen Wills Neuroscience Institute at UC Berkeley, aims to develop a novel brain-machine interface that uses light to monitor and modulate the activity of thousands to millions of individual neurons in the cerebral cortex.
Divisions
- Science Programs
Secondary Affiliation:
Environmental Genomics and Systems Biology
- Comparative and Functional Genomics
Recent Publications
Related News
Taking Stock of the Known and Unknown Microbial Space
In Science Advances, JGI researchers have taken stock of the current state of microbial genomic biodiversity. Using publicly available genome sequence data generated over the past three decades, their study assesses what fraction of the microbial diversity we know about, and proposes a path forward to curate and cultivate what is still unknown.
Doubling Down on Known Protein Families
Through a novel approach detailed in Nature, a massive computational analysis of microbiome datasets by the JGI focuses on unveiling protein functional diversity.
JGI Adds Actinobacteria Chapter in the Genomic Encyclopedia of Bacteria and Archaea
Large-scale comparative analysis leads to identification of biosynthetic gene clusters for novel secondary metabolites for multiple applications
Biography
Gary started his scientific career in 1978 as a research technician in the Schubiger lab at the University of Washington, where he worked on imaginal disc formation in Drosophila. He obtained his PhD in the lab of Charles Laird at the University of Washington, where he worked on the role of nucleolus organization in rDNA function. He finished his PhD in Genetics in 1987 and continued to work with Drosophila during his postdoc in the lab of Allan Spradling at the Carnegie Institute.
His work in the Spradling lab spurred his interest in heterochromatin formation and 4 years later he started his own lab at The Salk Institute, La Jolla in 1991. After being in San Diego for 12 years, during which he obtained his professorship at The Salk Institute for Biological Studies, Gary and his lab moved to the Lawrence Berkeley National Lab (LBNL) in 2003. Gary has been an adjunct professor at UC Berkeley since 2003 and became director of the LBNL Life Sciences Division in 2011.
The Karpen lab has a long-standing interest in chromatin structure and function, with a special emphasis on heterochromatic DNA regions. The current projects in the lab range from centromere formation and function, to the role of lncRNAs, ageing, and DNA repair in heterochromatin formation and maintenance.
Research Interests
Our studies are focused on understanding inheritance, chromatin structure, gene expression, and the organization of chromosomes in the nucleus. Most of our studies have focused on the fruit fly Drosophila melanogaster as a model for chromosome function in metazoans, which allows us to address mechanisms in animals by synergistically combining molecular, genetic, cell biological and biochemical approaches. Additionally, we have examined the relevance of our findings to human chromosomes, and have demonstrated surprising similarities between these evolutionarily-distant species.
Recent Publications
Related News
Secrets of the Centromere Revealed by First Gapless Human Genome Sequence
A years-long project has illuminated the sequences at the chromosomes’ center, where proteins bind to move replicated DNA into daughter cells. The newly completed genome, dubbed T2T-CHM13, represents a major upgrade from the current reference genome, called GRCh38, which is used by doctors when searching for mutations linked to disease, as well as by scientists looking at the evolution of human genetic variation.
Exploring Human Origins in the Uncharted Territory of Our Chromosomes
A group of geneticists from Berkeley Lab, UC Davis, UC Santa Cruz, and UC Berkeley are unraveling new details about human evolution by studying the uniquely regulated portion of our chromosomes that surround the centromeres. These stretches of DNA – termed centromere-proximal regions (CPRs) – are largely composed of highly repetitive, mostly non-gene-coding sequences that […]
Epigenetic Effects of ‘Genomic Parasites’ Impact Their Evolution
In a study published in eLife, Biological Systems and Engineering (BSE) postdoctoral researcher Grace Lee and senior scientist Gary Karpen investigated the extent to which transposons—bits of DNA that copy themselves and jump to other locations in the genome—harm organisms through epigenetic means, such as changing the way DNA is packaged in cells, and whether this influences how transposons evolve. In a Q&A with the journal, Lee explained the background of the research, the specific question she and Karpen were interested in, and the most illuminating result among their findings.
Research Interests
The Katz group applies emerging principles of supramolecular chemistry on solid surfaces towards the synthesis of functional advanced materials, involving catalysis and adsorption. We lead in the synthesis of organic-inorganic active sites, which are at the state-of-the-art in structural control. Our modular and generalizable engineering approach aims to roughly mimic principles of active site connectivity found in biological catalysts.
This approach leads to the highly refined design of heterogeneous catalysts, which function with an activity and selectivity that cannot be achieved using conventional approaches.
The group has most recently invented approaches for (i) mild zeolite precursor delamination, preseving framework cystallinity for applications benefiting from crystalline supports, (ii) the hydrolysis of glycosidic bonds under the mildest (i.e. close to pH 7) pH conditions that have ever been achieved, and (iii) design and synthesis of robust metal clusters that consist of an unprecedented amount of catalytic surface. Area (i) impacts catalytic applications where zeolitic supports are beneficial over traditional amorhous ones (see Grosso-Giordano et al. JACS 2018, 140, 4956—4960). Area (ii) broadly impacts the depolymerization and pretreatment of biomass to biofuels using less harsh chemicals. Area (iii) improves on the major limitation to cluster catalysts in industry—lack of stability (see Gates et al. Nature 1994, 372, 346—348).











