Among the 30 proposals selected for the 2018 Community Science Program (CSP) of the U.S. Department of Energy Joint Genome Institute (DOE JGI) is one by Jillian Banfield, a UC Berkeley professor and Berkeley Lab faculty scientist affiliated with the Environmental Genomics & System Biology (EGSB) Division. Banfield will study the impact of soil microbial communities along the East River in Colorado on the quality of water leaving the watershed. The research is part of a major DOE project to investigate how terrestrial systems respond to perturbations such as droughts, floods, and early snowmelt. The full list of accepted proposals can be viewed here.
American Society for Microbiology Honors Nikos Kyrpides
Nikos Kyrpides, head of the Prokaryote Super Program at the Department of Energy Joint Genome Institute (DOE JGI) and a senior scientist in the Biosciences Environmental Genomics and Systems Biology (EGSB) division, was selected as the 2018 USFCC/J. Roger Porter Award recipient by the American Society for Microbiology (ASM). He will be honored at the 2018 ASM Microbe Meeting in Atlanta, Ga. The award recognizes outstanding effort by a scientist in demonstrating the importance of microbial biodiversity through sustained curation or stewardship of a major resource used by the scientific community. For more than a decade, Kyrpides and his colleagues have been working toward the goal of developing a comprehensive catalog of reference genomes for every bacterial and archaeal species. Read more from JGI.
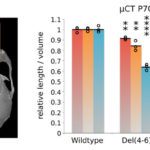
Modular Ensembles of Enhancers Regulate Ihh, Developmental Gene Expression in Mice
Biosciences researchers Axel Visel (JGI/EGSB) and Marco Osterwalder (EGSB) contributed to a study, led by researchers at the Max Planck Institute for Molecular Genetics, in which targeted genome editing and a transgenic reporter assay were used to characterize elements regulating Ihh (encoding Indian hedgehog) in mice. Indian hedgehog is a mammalian signaling protein involved with the development and proliferation of cells in cartilage. In humans, copy number variants (CNVs) upstream of Ihh cause localized phenotypes including premature fusion of the sutures of the skull and malformation of the phalanges. The study, published in Nature Genetics, showed that in mice Ihh is regulated by modular ensembles of enhancers (with individual tissue specificities) that appear to act in an additive manner. Despite apparent redundancy and overlapping function of enhancers, these ensembles—in which the correct number of each enhancer is present—are necessary for precise spatiotemporal control of developmental gene expression.
Another Successful Year for Biotech Partners Interns at Biosciences Area
The Biosciences Area partnered once again with Biotech Partners to provide paid summer internships to high school students. This year six high school students worked side by side with Biosciences researchers across the Area’s laboratories. The mission of the non-profit Biotech Partners is to educate underserved youth in the Bay Area with personal, academic and professional development experiences that increase participation in higher education and access to fulfilling science careers.
Council on Competitiveness Releases Report on Advancing U.S. Bioscience
Biosciences’ ALD Mary Maxon and Chief Science and Technology Officer Jay Keasling recently attended an event in Washington, D.C., sponsored by the Council on Competitiveness. The briefing on Capitol Hill marked the release of the council’s report, Leverage: Advancing U.S. Bioscience, which captures the outputs of a day-long discussion last year about infrastructure, technology, investment, and talent needed for advancements in U.S. bioscience.
- « Previous Page
- 1
- …
- 34
- 35
- 36
- 37
- 38
- …
- 46
- Next Page »
Was this page useful?