The Lab’s Computing Sciences and Biosciences Areas jointly held a week-long workshop focused on machine learning in data science, the goal of which was to build bridges through a common foundation in statistical computing. Thirty attendees from Biosciences, Computing Sciences, and Lab IT received hands-on training provided by Data Incubator before engaging in a “hackathon” to apply the techniques to relevant science problems and data sets.
Women Scientists & Engineers Council Marks 10th Anniversary
 Marking its 10th anniversary this year, the Women Scientists & Engineers Council (WSEC) is about to embark on a new set of initiatives to make the Lab an even better place to work, from partnering with HR on the development of new training opportunities, to exploring ways to improve communication around maternity leave for new hires.
Marking its 10th anniversary this year, the Women Scientists & Engineers Council (WSEC) is about to embark on a new set of initiatives to make the Lab an even better place to work, from partnering with HR on the development of new training opportunities, to exploring ways to improve communication around maternity leave for new hires.
Several Biosciences staff members are active in WSEC namely MBIB’s Jill Fuss, Work-Life Balance subcommittee, JGI’s Esther Singer, Empowerment subcommittee co-chair Esther Singer and Biosciences’ Astrid Terry, Networking subcommittee co-chair.
Read more in Today At Berkeley Lab.
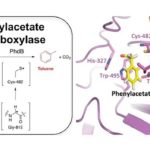
JBEI Enzyme Discovery Enables First-Time Microbial Production of the Octane Booster Toluene
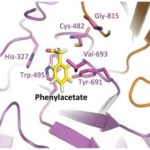 Researchers at the Department of Energy’s Joint BioEnergy Institute (JBEI) and Lawrence Berkeley National Laboratory (Berkeley Lab) have discovered a new enzyme that will enable microbial production of a renewable alternative to petroleum-based toluene, a widely used octane booster in gasoline that has a global market of twenty nine million tons per year.
Researchers at the Department of Energy’s Joint BioEnergy Institute (JBEI) and Lawrence Berkeley National Laboratory (Berkeley Lab) have discovered a new enzyme that will enable microbial production of a renewable alternative to petroleum-based toluene, a widely used octane booster in gasoline that has a global market of twenty nine million tons per year.
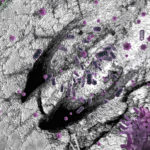
Plants Really Do Feed Their Friends
Researchers at Berkeley Lab and UC Berkeley have discovered that as plants develop, they craft their root microbiome, favoring microbes that consume very specific metabolites. Their study could help scientists identify ways to enhance the soil microbiome for improved carbon storage and plant productivity.
JGI Develops a Reference Catalog for the Rumen Microbiome
Reported March 19, 2018, in Nature Biotechnology, an international team including JGI scientists presents one of the largest targeted cultivation and sequencing projects to date: a reference catalog of rumen microbial genomes and isolates cultivated and sequenced from the Hungate1000 collection. The catalog was produced through the coordinated efforts of rumen microbiology researchers worldwide. At the beginning of the project, there were only reference genomes for 14 bacteria and one methanogen. The Hungate catalog now contains a total of 501 genomes—410 newly generated from this study, plus an additional 91 already publicly available from other studies. Read the full story on the JGI website.
- « Previous Page
- 1
- …
- 30
- 31
- 32
- 33
- 34
- …
- 46
- Next Page »
Was this page useful?