Deepanwita Banerjee
Computational Biologist Research Scientist
Research Interests
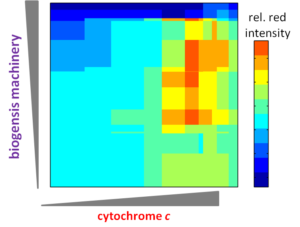
Deepanwita Banerjee, PhD, is currently working on computationally driven host engineering for growth-coupled production of biofuels and bioproducts using the Product Substrate Pairing (PSP) approach. She uses computational strain optimization methods and genome scale metabolic models (GSM) to predict gene targets for engineering Pseudomonas putida. She is also working towards predicting PSP strategies in P. putida for utilization of lignocellulose derived carbon sources for bioenergy and biomanufacturing applications. The goal is to test and learn to better design robust and sustainable microbial hosts that maintain desirable phenotype across industrially relevant scales and conditions.
Deepanwita Banerjee’s research interests include understanding microbial systems for desirable phenotypes using multi-omics integrated systems biology approach. The applications are diverse ranging from improving production of value-added products or assessing microenvironments for a redox imbalance in vivo. She is also interested in understanding the emergent properties of a microbial hosts that are a result of the complex relationship between metabolism and transcriptional regulation. Currently there are digital resources available for model microbes that help inform such relationships but is lacking for non-model microbial systems. Genome-scale network reconstruction of a microbial system requires integration of more than one layer of biological networks (e.g., metabolism, regulation or signaling) as constraints to improve prediction power of such computational models for desirable phenotypes such as growth coupled production or maximum bioconversion of carbon sources.
Programs & Initiatives
Recent Publications
Related News
Speeding up Biomanufacturing with a Turnkey Framework
Researchers from the Joint BioEnergy Institute (JBEI) developed a new framework that reduces the time of developing novel bioproducts. This new workflow, called Product Substrate Pairing (PSP), has already shown great promise for engineering strains that can convert common bacterial food sources into target molecules.
Deepanwita Banerjee, Multi-faceted Modeler
Deepanwita Banerjee learned the value of reusing things at an early age. This perspective continues to influence her life: from her DIY dollhouse project to her research that's recycling genes to build more sustainable products.
Microbe “Rewiring” Technique Promises a Boom in Biomanufacturing
A new approach to modifying microbes’ metabolic processes will speed up production of innovative bio-based fuels, materials, and chemicals Researchers from Lawrence Berkeley National Laboratory (Berkeley Lab) have achieved unprecedented success in modifying a microbe to efficiently produce a compound of interest using a computational model and CRISPR-based gene editing.
Divisions
- Science Programs
Secondary Affiliation:
Environmental Genomics and Systems Biology
- Comparative and Functional Genomics
Biography
Stephen Mondo completed his PhD training at Cornell University in 2013, where he studied host-endosymbiont interactions with Dr. Teresa Pawlowska. After a brief postdoc in the Bogdanove lab at Cornell studying TAL effectors present in fungal endosymbiont he joined the Fungal and Algal Genomics team at the DOE Joint Genome Institute as a Data Scientist. Here, Stephen supports the fungal scientific community through annotation of fungal genomes, comparative genomic analyses for user-driven publications and tool development for improved quality of fungal genome annotations. Stephen also conducts his own independent research in support of DOE mission objectives, primarily in the areas of epigenomics an chromatin regulation. Additionally, he now leads multi-omics data analysis, integration and visualization efforts within the Fungal and Algal Genomics program.
Research Interests
Epigenetics, chromatin regulation, fungal comparative genomics, evolutionary & co-evolutionary theory, host-endosymbiont interactions, phylogenomics, early-diverging fungi
Recent Publications
Related News
Biosciences FY25 LDRD Projects
The projects of 23 Biosciences Area scientists and engineers received funding through the FY25 Laboratory Directed Research and Development (LDRD) program.
Biosciences Area FY24 LDRD Projects
The projects of 21 Biosciences Area scientists and engineers received funding through the FY24 Laboratory Directed Research and Development (LDRD) program.
Biography
From the Molecular Foundry page:
Dr. Ajo-Franklin has been a Staff Scientist at the Molecular Foundry since 2007. Before that, she received her Ph.D. in Chemistry from Stanford University with Prof. Steve Boxer and was a post-doctoral fellow with Prof. Pam Silver in the Department of Systems Biology at Harvard Medical School.
Dr. Ajo-Franklin is fascinated by the incredible, diverse functionality of biological molecules and nanoassemblies, and seeks to engineer these complexes and their host organisms to address global challenges in energy and the environment.
Research Interests
Dr. Ajo-Franklin’s research creates engineered bioassemblies–electron nanoconduits–that direct the flow of charge between living cells and non-living materials. Though a combinatorial approach, she has established an unparalleled ability to genetically program the enormously complex biosynthesis of these electron nanoconduits and to quantitatively characterize their function at the biological-inorganic nanointerface. These capabilities have allowed her to discover design principles that govern the emergent processes of nanoconduit assembly and function. In future work, she will expand the combinatorial space through designed genetic variants and adaptive evolution approaches to dissect fundamental controls on charge transfer at this hybrid nanointerface.
ENGINEERING ELECTRONIC COMMUNICATION
Cellular-electrical connections would enable devices to combine specialties of the living world and the non-living technological world. This new class of smart, self-renewing nanostructured systems has the potential to revolutionize environmental sensing and energy harvesting applications and to open new avenues to program cellular behavior. Building towards this vision, the long-term objectives of the our research are to develop understanding of the principles which govern electron flow across living-non-living interfaces and to use those principles to create electron transfer pathways between individual living microbial cells and non-living electrodes. Our overall strategy is both to explore how naturally-occurring microbes achieve electron transfer to inorganic surfaces and to use genetic and materials surface engineering to create new ‘domesticated’ hybrid cell-electrode systems.
ASSEMBLY & APPLICATIONS OF S-LAYER PROTEINS
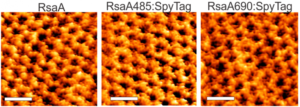
S (`surface’)-layer proteins form crystalline lattices on the outsides of many bacteria and archaea. These nearly ubquitous structures play a number of crucial biological roles: they serve as structural scaffolds, effect selective transport of ions and proteins, serve as templates for mineralization, and protect against phagocytosis. While the lattice structures of many S-layers are known, the dynamical mechanisms through which they form are poorly understood. A molecular-level picture of the assembly of the S-layer protein SbpA from Lysinibacillus sphaericus on lipid bilayers was obtained only recently by using in situ atomic force microscopy. AFM images elucidated the phase transition of amorphous precursors into the crystalline clusters composed of folded tetramers and subsequent growth without disappearance of any crystal clusters. Such a system demonstrates the non-classical crystallization behaviors in the S-layer assembly. Molecular dynamic simulations of the S-layer assembly using a coarse grain model also supported the non-classical nucleation arising from a dense liquid precursor, and subsequent growth of the crystal2. The long-term objectives of our research are to develop predictive understanding of S-layer protein nucleation and growth, so as to advance basic knowledge of the dynamics of coupled nanoscale phase separation and self-assembly and to enable control of the nanoscale structures. Together with the Whitelam, DeYoreo, and Bertozzi groups, we are using a combination of fluorescence microscopy, atomic force microscopy, and computer modeling to reveal the mechanisms underlying S-layer crystallization and to identify strategies for its control.
Recent Publications
Related News
Engineering Living ‘Scaffolds’ for Building Materials
Taking their cue from Nature, Berkeley Lab researchers have engineered living cells to act as a starting point, or scaffold, for the self-assembly of composite materials. The resulting engineered living materials (ELMs) represent a new class of material that may open the door to advanced applications in bioelectronics, biosensing, and smart materials. Leading the effort was Caroline Ajo-Franklin, whose lab is part of the Molecular Foundry, a DOE Office of Science User Facility, and who holds a secondary appointment in the Molecular Biophysics and Integrated Bioimaging (MBIB) Division. A study describing the work was recently published in ACS Synthetic Biology.
Gut Bacteria’s Shocking Secret: They Produce Electricity
UC Berkeley scientists have discovered that a common diarrhea-causing bacterium, Listeria monocytogenes, produces electricity using an entirely different technique from known electrogenic bacteria—and that hundreds of other bacterial species use this same process. The scientists worked Caroline Ajo-Franklin, a staff scientist at the Molecular Foundry who has a secondary appointment in Molecular Biophysics and Integrated Bioimaging, on this research. Read more from the UC Berkeley News Center.
Biosciences and Energy Sciences Areas Host Biomaterials Workshop
On July 16-17, Biosciences, along with the Energy Sciences Area, hosted an internal workshop to discuss and identify paths forward for biomaterials research at Berkeley Lab and in larger contexts–across the national laboratory complex, at universities, and in the private sector– that would enable the creation of new materials with performance-advantaged properties. This workshop continues the program development efforts established through the recent Advanced Biogenic Chemicals and Materials Laboratory-Directed Research and Development initiative established at Berkeley Lab in 2017.
Divisions
Molecular Foundry
Secondary Affiliation:
Molecular Biophysics and Integrated Bioimaging
- Cellular and Tissue Imaging
Biography
Bruce Cohen, Ph.D. is a Staff Scientist at the Molecular Foundry and the Division of Molecular Biophysics & Integrated Bioimaging at Lawrence Berkeley National Laboratory in Berkeley, California (USA), where his research focuses on the design and synthesis of novel luminescent nanomaterials for bioimaging. He graduated with honors in Chemistry from Princeton and earned his Ph.D. in Chemistry and Biophysics from UC Berkeley working with Daniel E. Koshland, Jr. He earned a certificate in Neurobiology from Woods Hole Institute before working as a Howard Hughes Medical Association postdoctoral fellow in the laboratory of Lily Y. Jan (UCSF) developing organic and protein-based fluorescent probes for addressing problems in protein biophysics and bioimaging. He has a broad background covering nanoscience, biophysics, neuroscience, photophysics, and chemical biology. His current research interests include development of novel organic fluorophores for imaging and sensing, nanoparticle-based biosensors, and lanthanide-based upconverting nanoparticles as single molecule, quantitative, and deep tissue imaging probes.
Research Interests
Optical microscopy is the primary means of studying live cells in real time, enabling analysis of individual cellular components at high spatial and temporal resolution. Inorganic nanocrystals have shown promise as transformative probes for these imaging studies, with exceptional optical properties not found in other materials. Along these lines, we are developing novel nanocrystals as biosensors and single-molecule probes, improving bioconjugation and targeting chemistries, and imaging live cells with these reagents. We aim to integrate the development of novel luminescent nanomaterials into multidisciplinary efforts to address significant biological questions of cell function.
Imaging single molecules
Lanthanide-based upconverting nanoparticles (UCNPs) sum the energies of 2 incident NIR photons to emit one at visible wavelengths, an unusual property unlike anything found in the cell. UCNPs have significant advantages over other luminescent reporters, including an absence of on–off blinking, single-molecule multiphoton NIR excitation at powers approaching those used for standard one-photon confocal imaging, no overlap with cellular autofluorescence, and no measurable photobleaching under prolonged single-particle excitation. Our synthetic efforts have established control over UCNP size to produce smaller nanocrystals more compatible with many imaging applications. Current efforts are aimed at optimizing single nanocrystal brightness for extended single-molecule imaging in live cells, and developing UCNPs capable of chemical sensing for studying cellular biochemistry.
Optical biosensors
Many nanocrystals exhibit brightness and stability far superior to conventional fluorophores, making them ideal as the bases for biosensing reagents. We have developed quantum dot and UCNP energy transfer-based systems for the sensitive detection of cellular chemistry and protein motions. Our current work focuses on developing bright, selective sensors for studying neuronal activity, including voltage sensors, sensors of neurotransmission in the brain, and ion sensors.
Nanocrystal biocompatibility and targeting
Broadening the scope of nanocrystal surface conjugation chemistry is essential to expand their reach for imaging applications. High-quality UCNPs and quantum dots are synthesized in hydrophobic solvent and must be transferred to water and made biocompatible to have any utility as imaging probes or biosensors. An ongoing challenge in applying nanocrystals as probes for cellular imaging is improving their aqueous passivation and developing new reactions that work on nanocrystal surfaces. A growing area of research for us is development of new bioconjugation chemistries, including click reactions, SpyCatcher ligation, and bioorthonganol reactions for immunotargeting.
Recent Publications
Related News
Congratulations to Biosciences Area Director’s Award Recipients
Each year, the Berkeley Lab Director’s Achievement Award program recognizes outstanding contributions by employees to all aspects of Lab activities. Several Biosciences Area personnel are among the 2025 honorees.
Shine On: Avalanching Nanoparticles Break Barriers to Imaging Cells in Real Time
The diffraction limit is a fundamental property of light that has long prevented optical microscopes from bringing into focus anything smaller than half the wavelength of visible light (~200 nanometers), which is at least an order of magnitude larger than the tiny protein machines that keep cells, and us, running. A team of researchers co-led scientists in Berkeley Lab's Molecular Foundry and Columbia University’s school of engineering developed a new class of crystalline material that, when used as a microscopic probe, overcomes the diffraction limit without heavy computation or a super-resolution microscope. The amazing new material, called avalanching nanoparticles (ANPs), will advance high-resolution, real-time bio-imaging of a cell’s organelles and proteins, as well as the development of ultrasensitive optical sensors and neuromorphic computing that mimics the neural structure of the human brain, among other applications. The work was reported in a cover article in the journal Nature.
Divisions
- Science Programs
Secondary Affiliation:
Environmental Genomics and Systems Biology
- Comparative and Functional Genomics
Biography
I received my B.S. in Bioinformatics and Molecular Biology at Rensselaer Polytechnic Institute in 2006. From there, I went on to pursue a Ph.D in Biological Sciences at UCSD, where I trained in the laboratory of Dr. Joanne Chory at the Salk Institute, studying morphometric features associated with shade avoidance in Arabidopsis thaliana. I then undertook a postdoctoral position in Dr. Steve Kay’s laboratory at UCSD to study circadian and diurnal growth and transcriptome profiles of the model grass, Brachypodium distachyon, before taking on a second postdoctoral position at the DOE-Joint Genome Institute using genome-wide mutagenesis surveys to investigate root colonization by soil bacteria. I became a Research Scientist at LBNL in 2020, and received a DOE Early Career Award in 2021.
Research Interests
Plants are phenomenal in their ability to adapt to adverse conditions, harboring an incredible diversity of responses to environmental stress. I am broadly interested in studying exactly how plants interact with their environment, from their relationships with soil micoorganisms to their reactions to drought, light, and nutrient scarcity. At the Joint Genome Institute, my research focus has been on understanding how microbes can colonize plant roots, focusing on genetic components required for effective colonization. In addition, I am now focusing on how individual plant cells respond to their biotic and abiotic environment using emerging single-cell characterization technologies (scRNA-seq and Spatial Transcriptomics).
Recent Publications
Related News
How Plants and Mycorrhizal Fungi Cooperate
Researchers have studied both sides of plant-fungi symbiosis in one of the first cross-kingdom spatially-resolved transcriptomics studies to date.
Report from Second Plant Single-cell Solutions for Energy and the Environment Workshop Available
On April 29, 2021, Berkeley Lab hosted a second workshop to identify the most pressing barriers to wider adoption of single-cell sequencing and omics technologies, and to discuss solutions to remedy those barriers in order to drive discovery. The workshop report is now available for download.
Cole Named DOE Early Career Awardee
The JGI's Ben Cole is one of five Berkeley Lab scientists selected by the U.S. Department of Energy’s Office of Science to receive funding through the Early Career Research Program (ECRP). Under the program, researchers based at DOE national laboratories will receive $500,000 per year, for five years, to cover salary and research expenses. His award is for a project that will employ sequencing and molecular profiling techniques to examine the genes and gene-regulating processes underlying how individual cells in two prominent bioenergy crops – sorghum and switchgrass – respond to drought and nutrient limitation.
Divisions
- Science Programs
Secondary Affiliation:
Environmental Genomics and Systems Biology
- Biosystems Data Science
Research Interests
- Annotation of fungal genomes to better understand their roles in interactions with e.g. plants, algae, and other fungi
- Comparative phylogenomics to help resolve the Fungal Tree of Life, particularly untangling the lesser studied basal fungal clades
- Elucidation of the underlying biological mechanisms of efficient biomass degradation by fungi
Recent Publications
Related News
JGI Researchers Trace the Evolution of Shiitake Mushrooms
By complete sequencing of 24 new mushroom genomes, and assembling genomes from 60 existing sequences, this work expands the family tree for Lentinula. Within those samples, this study also tracks the enzymes these fungi use to break down wood, and compares genetic diversity between cultivated and wild shiitake populations.
JGI Develops Single-Cell Pipeline for Fungal Diversity
More than a million species of fungi are estimated to live on this planet, but most of that diversity remains unknown because the fungi have avoided detection and have not been cultured for study in laboratories. A team led by researchers at the Joint Genome Institute has developed a pipeline to generate genomes from single cells of uncultivated fungi. The approach was tested on several uncultivated fungal species representing the earliest evolutionary branches in the fungal genealogy that provide a repertoire of important and valuable gene products.
Divisions
- Science Programs
Secondary Affiliation:
Environmental Genomics and Systems Biology
- Comparative and Functional Genomics
Research Interests
- Comparative analysis of fungal and algal genomes
- Annotation and prediction of gene function
- Integration of diverse biological data
Recent Publications
Related News
Biosciences Area FY23 LDRD Projects
22 Biosciences Area scientists and engineers were awarded funding for their projects through the FY23 Laboratory Directed Research and Development (LDRD) program.

Building: 91, Room 210c
Mail Stop: 91R0183
Phone: 510-495-8511
ikblaby@lbl.gov
https://jgi.doe.gov/our-science/scientists-jgi/ian-blaby/
Links
Divisions
Secondary Affiliation:
Environmental Genomics and Systems Biology
- Comparative and Functional Genomics
Research Interests
Dr. Blaby joined the JGI in 2019 as the lead of the DNA synthesis platform, where he manages production for approved user projects as well as leading computational and lab-based R&D efforts for synthetic biology and functional genomics. Prior to JGI, he was a group lead at Brookhaven National Laboratory where he focused on functional genomics of phototrophs. Through this and post-doctoral positions at University of Florida and UCLA he has worked with a wide range of eukaryotic algae, archaea and bacteria with a view to developing a deeper understanding of sequence-based gene function.
Recent Publications
Related News
Studying the Tiniest Archaea, JGI Users Find a Genomic Switch of Friend or Foe
Meta-omics datasets show that CRISPR-Cas systems determine mutualism or parasitism between some archaeal hosts and their hitchhikers.
Leveraging Soil Virus for Insight into Maintaining Microorganisms
Using data generated from the JGI’s IMG/M Data Portal, researchers have ID’d a protein within soil virus that could contribute to carbon cycling.
Onsite PhD Student Visit Amps Up Collaborative Spirit
Biosciences Area staff recently welcomed 40 PhD students from Wageningen University in the Netherlands. Over two days, they hosted the contingent at Emery Station East (ESE) and the Integrative Genomics Building (IGB).

Building: 977, Room 0267
Mail Stop: M/S 977
Phone: 510-486-7069
KMChristiansen@lbl.gov
https://biosciences.lbl.gov/program-development-group/
Biography
As Area Deputy, Katy Christiansen supports Biosciences ALD Paul Adams, helping to plan and execute long-range strategies to advance the Area’s scientific mission. After earning her PhD in plant science from Indiana University, Bloomington, Christiansen first joined Berkeley Lab in 2008 as a postdoctoral researcher in the plant systems biology group at JBEI. After two years as a AAAS Science & Technology Policy Fellow at the DOE Bioenergy Technologies Office, where she served as a technical adviser and assisted in development of funding opportunities, she returned to the Biosciences Area in 2014 to help lead strategic planning efforts. Her efforts resulted in direct funding from DOE that established the multimillion dollar Agile BioFoundry (ABF) and Trial Ecosystem Advancement for Microbiome Science (TEAMS) programs. As Head of the Biosciences Program Development Group (BPDG) since its inception in 2020, she now leads program development activities and strategic planning for the Area, including building new multi-institutional research programs with other national laboratories. Christiansen is also responsible for the Biosciences Strategic Plan (BSP), a 10-year scientific strategic plan that describes biosciences research aspirations in energy, environment, health, biomanufacturing, and technology development.
Related News
Biological Systems and Engineering Division and Program Leadership Changes
Division Director Blake Simmons announced that, effective March 2, Chris Petzold will lead the Biodesign Department as Interim Head following Nathan Hillson’s departure. Petzold will also assume the role of Chief Information Officer for the Joint BioEnergy Institute, as announced by CEO Jay Keasling. Katy Christiansen will serve as the lead principal investigator of the Agile BioFoundry.
Katy Christiansen Named Biosciences Area Deputy for Science
ALD for Biosciences Paul Adams has appointed Katy Christiansen to the role of Area Deputy for Science following a national search. Christiansen, who has served as Interim Area Deputy since 2021, will work with Adams to support the Biosciences Area’s scientific mission and be responsible for helping to plan and execute our long-range strategy.
Report from Second Plant Single-cell Solutions for Energy and the Environment Workshop Available
On April 29, 2021, Berkeley Lab hosted a second workshop to identify the most pressing barriers to wider adoption of single-cell sequencing and omics technologies, and to discuss solutions to remedy those barriers in order to drive discovery. The workshop report is now available for download.
Research Interests
Dr. Justin Reese is a Research Scientist in the Environmental Genomics and Systems Biology Division at Lawrence Berkeley National Laboratory. His research seeks to understand how complex biological systems give rise to health and disease, and how diverse data can be harnessed to accelerate biological discovery. He is particularly interested in questions that bridge biology and biomedicine, with an emphasis on translating biological knowledge into tools that improve human health.
He develops and applies computational approaches that include artificial intelligence, machine learning, knowledge graphs, and ontologies. His work spans molecular, clinical, and environmental biosciences. He leads and co-leads multi-institutional collaborations that combine structured biological knowledge with modern AI systems to derive insights from complex biological datasets. His projects include constructing large-scale knowledge graphs and applying graph machine learning algorithms to them, using large language models to extract and organize information from scientific literature and clinical data, and developing AI systems that assist scientists in discovery, including applications such as improving rare disease diagnosis, using AI to accelerate research in neurodegenerative diseases, and building tools to suggest better candidates for drug repurposing.
Recent Publications
Related News
Machine Learning Tackles Long COVID
A new machine learning tool developed by a team of researchers led by Justin Reese of Berkeley Lab and Peter Robinson of Jackson Lab analyzes electronic health records to find symptoms in common between people who have been diagnosed with long COVID and to define subtypes of the condition.
Metformin May Mitigate More Severe COVID-19 Outcomes in Patients with Prediabetes, PCOS
An international team led by Justin Reese, a research scientist in the Environmental Genomics and Systems Biology (EGSB) Division, analyzed electronic health record data aggregated in the National COVID Cohort Collaborative (N3C) Data Enclave to assess whether metformin is associated with reduced COVID-19 severity in people with prediabetes or polycystic ovary syndrome (PCOS), two common conditions that increase the risk of severe COVID-19 presentation.
Research Interests
High-performance computing
Anomalous diffraction
XFEL diffraction
Chemical crystallography
Recent Publications
Related News
Congratulations to Biosciences Area Director’s Award Recipients
Several Biosciences Area personnel are among the 2024 recipients of Berkeley Lab Director’s Achievement Awards. The program recognizes outstanding contributions by employees to all aspects of Lab activities.
Researchers Capture Elusive Missing Step in Photosynthesis
After decades of effort, scientists have revealed atomic-scale details of the water splitting step of photosynthesis, the chemical process that generates the air we breathe. The latest work adds to our understanding of photosynthesis and will aid the development of fully renewable alternative energy sources.
Crystallography for the Misfit Crystals
A team of researchers is working to provide a better way for scientists to study the structures of the many materials that don’t form tidy single crystals. Their new technique, called small-molecule serial femtosecond X-ray crystallography, or smSFX, supercharges traditional crystallography with the addition of custom-built image processing algorithms and an X-ray free electron laser (XFEL).

Building: Integrative Genomics Building (IGB), Room 125
yezhangding@lbl.gov
http://www.northenlab.org/
Links
Research Interests
I am interested in using integrative system biology and high throughput approaches to understand plant specialized metabolism in plant environmental stress interactions. One of his projects is focused on plant microbiome interactions. Plants utilize chemicals to communicate with microorganisms, favoring beneficial microbes and killing harmful ones. I use integrative approaches, including metabolomics, genetics, transcriptomics, proteomics and others to identify novel secondary metabolites involved in plant microbe interactions and explore how plants use these metabolites to benefit their own.
Recent Publications
Related News
EcoFABs Could Help Fuel AI in Agriculture
A first-of-its-kind global study showed that EcoFABs can deliver consistent results across labs on three continents, supported by open protocols, tools, and datasets. The reliable, large-scale data EcoFABs generate are ideal for training AI, which could help accelerate discoveries in crop development, soil health, and agriculture.
EcoFAB: A Tool for Combating Climate Change and Training the Next Generation
Fabricated ecosystems—EcoFABs—are plastic, takeout box–sized growth chambers developed at Berkeley Lab to be a standardized and reproducible platform for conducting experiments on model plants and the microbes that live around their roots. A greater understanding of how plants and microbes work together to store vast amounts of atmospheric carbon in the soil will help in the design of better bioenergy crops for the fight against climate change.