Krishna K. Niyogi
Biologist Senior Faculty Scientist
Research Interests
My long-term research goals are to understand how photosynthetic energy conversion works, how it is regulated, and how it might be improved to help meet the world’s needs for food and fuel. We use a wide array of experimental organisms and interdisciplinary approaches to investigate fundamental questions about assembly, regulation, and dynamics of photosynthesis. Current lab members study the biosynthesis and function of photosynthetic pigments, assembly and repair of photosynthetic reaction centers, structure and dynamics of the photosynthetic membrane, mechanisms involved in sensing excess light, singlet oxygen signaling, transcriptional regulation of photosynthesis and photoprotection by light and carbon, and regulation of photosynthetic light harvesting in saturating light. By comparing how photosynthesis works in diverse organisms, we hope to uncover general design principles of natural photosynthesis as well as unique adaptations to different environments.
Recent Publications
Related News
How Algae Use Memory to Adapt to Sudden Changes in Sunlight
A new study co-led by Graham Fleming a senior faculty scientist in the Molecular Biophysics and Integrated Bioimaging (MBIB) Division, and Krishna Niyogi, a faculty scientist in MBIB, reveals the precise molecular machinery that underpins photoprotective memory in green algae. The results may help scientists develop more productive plants and improve crop yields.
Bioscientists to Receive DOE Funding for Biomanufacturing and Microbiome Research
Biosciences researchers are among the recipients of four new DOE awards. Two awards will focus on reducing carbon emissions while producing bioenergy. The other two are aimed at understanding the role of microbiomes in the biogeochemical cycling of elements like carbon.
Scientists Talk Diatoms in Genome Insider Podcast
Berkeley Lab scientists discuss their research on a tiny group of algae with insanely gorgeous exterior shells.
Divisions
Environmental Genomics and Systems Biology
- Molecular EcoSystems Biology
Secondary Affiliation:
Biography
Metabolomics Program Lead
Deputy Division Director, Environmental Genomics and Systems Biology
Senior Scientist, Berkeley Lab
Education
B.S. in Chemical Engineering, University of California Santa Barbara;
Ph.D. in Chemistry and Biochemistry, Arizona State University;
Post-doc in Metabolomics, The Scripps Research Institute;
Executive Education, UC Berkeley Haas School of Business;
DOE Oppenheimer Science and Energy Leadership
Summary
Dr. Trent Northen is currently a Senior Staff Scientist at Berkeley Lab. Since joining Berkeley lab in 2008 Dr. Northen has had leadership roles within large DOE funded research programs using mass spectrometry for energy and environmental research including through the Joint BioEnergy Institute and the ENIGMA SFA project. In 2015, he founded what is now the JGI Metabolomics Program to provide advanced metabolomic methods available to JGI users. Dr. Northen’s research is focused on using metabolomics to link genomes with environments to understand how webs of microbes cycle carbon and sustain biomes via the development and application of advanced mass spectrometry approaches and model laboratory ecosystems (EcoFABs).
Recent Awards and Service
2019 R&D100 Award, 2016 Berkeley Lab Director Exceptional Achievement Award, 2014 DOE Early Career Award, 2013 R&D100 award, 2009 Presidential Award for Science and Engineering PECASE awarded by President Obama, A JBEI Excellence Award, multiple LBNL SPOT Awards, and two NIH SPORE Career Development Awards. Dr. Northen currently serves as the Deputy Director of the Environmental Genomics and Systems Biology at Lawrence Berkeley National Laboratory, he was the interim Division Director within this same organization during the formation of the Division in 2015-2016. Dr. Northen regularly chairs workshops, organizes conference sessions, and serves on review panels. At Berkeley Lab he serves on a number of committees and is the Strategy Mentor for the Berkeley Lab Biosciences Environmental Strategy.
Research Interests
Please see our lab’s website for more detail on our research.
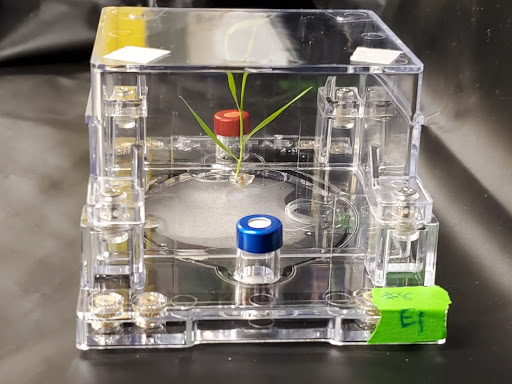
Summary: The combined impact of changing climate, population growth, environmental contamination, and soil degradation represent one of the greatest challenges facing humanity. Harnessing microbes will be a cornerstone of future sustainable agricultural and land management practices. Our laboratory uses mass spectrometry and fabricated ecosystem approaches to study the dynamic and reciprocal processes by which microbes interact to transform the organic molecules within their environment. Through the use of exometabolite profiling, we predict microbial metabolic webs in soils, sediment, and the rhizosphere for testing in fabricated ecosystems (EcoFABs) through the use of synthetic communities. Together these studies are providing insights into how microbes can be used to restore soil carbon, promote plant growth, and remove environmental contaminants.

Programs & Initiatives
Recent Publications
Related News
When Marine Algae Get Sick: How Viruses Shape Microbe Interactions
Researchers in the Environmental Genomics and Systems Biology Division collaborated on a study to better understand the role of viruses that infect photosynthetic phytoplankton in the marine food web.
Nurturing STEM Opportunities for Native Americans
A Berkeley Lab internship program aims to help increase Native American representation in graduate programs.
Introducing RhizoNet: AI-driven Plant Root Analysis
Berkeley Lab scientists from the Applied Mathematics and Computational Research (AMCR) and Environmental Genomics and Systems Biology (EGSB) Divisions developed RhizoNet, which harnesses the power of artificial intelligence (AI) to automate the process of root image analysis with exceptional accuracy.
Divisions
- Science Programs
Secondary Affiliation:
Environmental Genomics and Systems Biology
- Comparative and Functional Genomics
Recent Publications
Related News
JGI Researchers Trace Strawberry Ancestors For a View of Crop Evolution
Adam Session and Daniel S. Rokhsar have recently published a Nature Communications paper that builds off their early collaborations that could help crop breeders and researchers predict how crops and model organisms may evolve.
Congratulations 2021 Chan Zuckerberg Biohub Investigators
Four faculty scientists in the Biosciences Area were included in The Chan Zuckerberg Biohub Investigator Program, awarding $21 million to 21 University of California, Berkeley researchers.
Recent Publications
Related News
Pam Ronald Elected to American Academy of Arts and Sciences
Pam Ronald, a faculty scientist in the Environmental Genomics and Systems Biology (EGSB) Division, has been elected to the American Academy of Arts and Sciences. She was recognized for her accomplishments and leadership in plant pathology research.
Pamela Ronald Awarded 2022 Wolf Prize in Agriculture
Ronald was recognized for her work on disease resistance and environmental stress tolerance in rice.
JBEI’s Pamela Ronald Named GCHERA World Agriculture Prize Laureate
Pamela Ronald, the Joint BioEnergy Institute's (JBEI) Scientific Lead for Plant Pathology in the Feedstocks division, has been named the 2020 World Agriculture Prize laureate by the Global Confederation of Higher Education Associations for Agricultural and Life Sciences, or GCHERA. Ronald is a distinguished professor in the Department of Plant Pathology at the University of California, Davis (one of JBEI’s six academic research partners) and with the UC Davis Genome Center. She becomes the first woman whose work is recognized by the award. The virtual award ceremony will be held on November 30.
Research Interests
The Ryan lab studies mechanisms that regulate protein activity in bacteria. We have focused on the Gram-negative aquatic bacterium Caulobacter crescentus because it has a complex life cycle that is driven in large part by changes in the phosphorylation, proteolysis, localization, and interactions of proteins.
C. crescentus undergoes a developmental step during its division cycle from a motile cell unable to replicate its DNA to a nonmotile cell that is competent for chromosome replication. We use synchronized populations of motile G1-phase cells to study the events that occur during development. C. crescentus also divides asymmetrically, producing one motile and one nonmotile cell at each division. Proteins in the predivisional cell are segregated to opposite ends to mediate the differentiation of identity after division. We study the locations and movements of individual proteins using fluorescence microscopy.
Specific problems that we are investigating include the following: regulated degradation of a key transcription factor during the motile-sessile developmental step; the roles, locations, and interactions of enzymes that participate in cell wall biosynthesis; and the regulatory functions of protein phosphorylation on serine, threonine, or tyrosine residues.
Recent Publications

Building: 978, Room 4116
Mail Stop: 978-4121
Phone: (510) 486-7808
Fax: (510) 486-5225
BASimmons@lbl.gov
Biography
In addition to directing the Biological Systems & Engineering Division, Blake Simmons serves as the Chief Scientific and Technology Officer and Vice President of the Deconstruction Division at the Joint BioEnergy Institute in Emeryville. After earning his BS in chemical engineering from the University of Washington, Simmons continued his studies at Tulane University and received his doctorate in the same field. For the past 15 years, Simmons has been part of the Senior Management team at Sandia National Laboratories, most recently serving as the Senior Manager of Advanced Biomanufacturing, Biomass Program Manager, as well as Adjunct Professor at the University of Queensland. His expertise includes biofuels, renewable chemicals, biomanufacturing, abiotic-biotic interfaces, biomass pretreatment, enzyme engineering, biofuel cells, templated nanomaterials, microfluidics, desalination, and biomineralization.
Research Interests
Bioenergy
A significant focus in my research program at the Joint BioEnergy Institute is the development of new scientific insights and novel technologies for biomass deconstruction, the process of breaking down and fractionating lignocellulos into targeted intermediates through the use of ionic liquids. This includes the development of microbes and enzymes that can tolerate and operate efficiently in the presence of these ionic liquids, and the advent of consolidated conversion operations in order to produce a scalable, sustainable, and affordable technology suitable for deployment in a biorefinery.
Biotechnology and Biomanufacturing
While the 21st century will undoubtedly be recognized for advances in the biological and life sciences, another important element in magnifying the positive impacts of these advances is to enable the US bioeconomy, especially in the realms of developing advanced biomanufacturing technologies to replace and displace current manufacturing techniques that are energy intensive and have a negative effect on the environment. My goal is to mature the science of biotechnology and synthetic biology into a BioFoundry, and to identify opportunities where biomanufacturing can have a positive impact on cost and sustainability profiles relative to conventional manufacturing approaches.
Nanotechnology and Nanostructured Materials
Another significant research interest is the development of templated nanomaterials that provide a superior performance attributes relative to bulk materials. I have worked with surfactant-based approaches for the production of tailored nanostructures with unique properties as semiconductors, photocatalysts, catalysts, and energy storage materials.
Selected publications
- Sun, N., Xu, F., Sathitsuksanoh, N., Thompson, V.S., Cafferty, K., Li, C., Tanjore, D., Narani, A., Pray, T.R., Simmons, B.A., Singh, S. 2015. Blending municipal solid waste with corn stover for sugar production using ionic liquid process. Bioresour Technol, 186(0), 200-206.
- Cheng, G., Zhang, X., Simmons, B., Singh, S. 2015. Theory, practice and prospects of X-ray and neutron scattering for lignocellulosic biomass characterization: towards understanding biomass pretreatment. Energy Environ. Sci., 8(2), 436-455.
- Papa, G., Rodriguez, S., George, A., Schievano, A., Orzi, V., Sale, K.L., Singh, S., Adani, F., Simmons, B.A. 2015. Comparison of different pretreatments for the production of bioethanol and biomethane from corn stover and switchgrass. Bioresour Technol, 183C(0), 101-110.
- Socha, A.M., Parthasarathi, R., Shi, J., Pattathil, S., Whyte, D., Bergeron, M., George, A., Tran, K., Stavila, V., Venkatachalam, S., Hahn, M.G., Simmons, B.A., Singh, S. 2014. Efficient biomass pretreatment using ionic liquids derived from lignin and hemicellulose. Proc Natl Acad Sci U S A, 111(35), E3587-95.
- Eudes, A., Sathitsuksanoh, N., Baidoo, E.E., George, A., Liang, Y., Yang, F., Singh, S., Keasling, J.D., Simmons, B.A., Loque, D. 2015. Expression of a bacterial 3-dehydroshikimate dehydratase reduces lignin content and improves biomass saccharification efficiency. Plant Biotechnol J.
- Uppugundla, N., da Costa Sousa, L., Chundawat, S.P., Yu, X., Simmons, B., Singh, S., Gao, X., Kumar, R., Wyman, C.E., Dale, B.E., Balan, V. 2014. A comparative study of ethanol production using dilute acid, ionic liquid and AFEX pretreated corn stover. Biotechnol Biofuels, 7(1), 72.
- Sun, N., Parthasarathi, R., Socha, A.M., Shi, J., Zhang, S., Stavila, V., Sale, K.L., Simmons, B.A., Singh, S. 2014. Understanding pretreatment efficacy of four cholinium and imidazolium ionic liquids by chemistry and computation. Green Chemistry, 16(5), 2546-2557.
- Sathitsuksanoh, N., Holtman, K.M., Yelle, D.J., Morgan, T., Stavila, V., Pelton, J., Blanch, H., Simmons, B.A., George, A. 2014. Lignin fate and characterization during ionic liquid biomass pretreatment for renewable chemicals and fuels production. Green Chemistry, 16(3), 1236-1247.
- Ruegg, T.L., Kim, E.M., Simmons, B.A., Keasling, J.D., Singer, S.W., Soon Lee, T., Thelen, M.P. 2014. An auto-inducible mechanism for ionic liquid resistance in microbial biofuel production. Nat Commun, 5, 3490.
- Oleskowicz-Popiel, P., Klein-Marcuschamer, D., Simmons, B.A., Blanch, H.W. 2014. Lignocellulosic ethanol production without enzymes–technoeconomic analysis of ionic liquid pretreatment followed by acidolysis. Bioresour Technol, 158, 294-9.
Programs & Initiatives
Recent Publications
Related News
Two Scientists Named AAAS Fellows
Two scientists in the Biosciences Area, Abby Dernburg and Blake Simmons, have been elected into the 2025 class of the American Association for the Advancement of Science (AAAS).
Biological Systems and Engineering Division and Program Leadership Changes
Division Director Blake Simmons announced that, effective March 2, Chris Petzold will lead the Biodesign Department as Interim Head following Nathan Hillson’s departure. Petzold will also assume the role of Chief Information Officer for the Joint BioEnergy Institute, as announced by CEO Jay Keasling. Katy Christiansen will serve as the lead principal investigator of the Agile BioFoundry.
Breaking Through the Lignin Problem
A strategy developed at JBEI streamlines the process of converting lignin, a complex plant polymer, into starting materials for biomanufacturing processes.
Recent Publications
Related News
Using Bacteria to Accelerate Carbon Dioxide Capture in Oceans
Peter Agbo, a staff scientist in the Chemical Sciences Division, with a secondary appointment in the Molecular Biophysics and Integrated Bioimaging (MBIB) Division, has proposed a novel method for direct ocean capture of carbon using microbes. Removing CO2 from the oceans will enable them to continue to do their job of absorbing excess CO2 from the atmosphere.
Biosciences Area FY21 LDRD Projects
The projects of 15 Biosciences Area scientists and engineers received funding through the FY21 Laboratory Directed Research and Development (LDRD) program.
Biosciences Area FY19 LDRD Projects
The projects of 13 Biosciences Area scientists and engineers received funding through the FY19 Laboratory Directed Research and Development (LDRD) program. The funded projects span a diverse array of topics and approaches including the harnessing of microbiome data to uncover patterns of mutualism, evaluating radiobiological effects of laser-accelerated ion beams, improving bioenergy yield under drought stress, and the application of machine learning in tomogram segmentation. Lab-wide, 89 projects were selected from a field of 158 proposals. Biosciences Area efforts account for 15.07 percent of the $22.2 million allocated.
Biography
Antoine Snijders is the Division Leader of the Biosciences and Biotechnology (BBTD) Division in the Physical and Life Sciences Directorate. Antoine holds a Ph.D. in Cancer and Molecular Biology from Utrecht University and an M.S. in Biomedical Sciences from VU University Amsterdam (both in the Netherlands). After appointments as a Postgraduate Researcher and Postdoctoral Scholar at the Cancer Research Institute at the University of California, San Francisco, Antoine joined Lawrence Berkeley National Lab where he served as the Head of the Bioengineering and Biomedical Sciences Department. Antoine’s research focused on understanding the complex interactions among genetic background, environmental exposures, and the microbiome in determining disease risk.
Research Interests
My research goals are to understand cancer tumorigenesis specifically addressing key questions concerning the contribution of host genetics, environmental exposures and their interactions in cancer risk and tumor progression. My laboratory uses a systems biology approach, together with novel mouse models to identify genetic networks controlling susceptibility to cancer risk.
Precision medicine is an emerging approach for disease treatment and prevention that takes into account individual variability in environment, lifestyle and genes for each person. Genetic susceptibility is a major component that contributes to the variability in disease susceptibility. Thus, identifying the genes involved in susceptibility to cancer may have potential utility in developing novel personalized medicines, lead to greater understanding of the biological pathways involved in cancer development, and elucidate how environmental factors exert their effects in combination with genetic variants.
The broad and long-term goals of my laboratory are to identify how the interactions of combinations of genes and their functional polymorphisms and environmental exposures (including for example thirdhand smoke) contribute to disease susceptibility of individual human subjects. Major problems in assessing human cancer risk are that humans are genetically heterogeneous and exposures are pervasive and difficult to quantitate. Parallel exposures to multiple chemicals and lifestyle factors that can negatively impact health further confound efforts to assess risk.
Population-based mouse models offer many advantages for the study of the genetic basis of complex traits, including cancer, because of our ability to control both the genetic and environmental components of risk. My lab exploits the power of mouse genetics using Collaborative Cross (CC) mice, together with OMICS analyses to determine the influence of individual variations in disease susceptibility. This comprehensive systems biology approach will likely identify specific genes or pathways that are differentially controlled between mouse strains, and contribute to human variation in susceptibility to environment factor-induced carcinogenesis.
To learn more about thirdhand smoke, visit the Thirdhand Smoke Resource Center (www.thirdhandsmoke.org).
Recent Publications
Related News
A New Genetic Hallmark for Predicting Breast Cancer Outcomes
Biosciences researchers have used Collaborative Cross mice, a mouse model system designed to mimic the diversity of human populations, to identify a set of genetic factors that could help refine treatment approaches for a fast-growing form of breast cancer.
Biological Systems and Engineering Division Leadership Changes
Biological Systems and Engineering (BSE) Division Director Blake Simmons announced that following a search, Justin Heady has been named the BSE Deputy for Operations. There will also be changes in the leadership of the Bioengineering and Biomedical Sciences Department, effective August 12, 2024.
Infrared and AI Detect Low-dose Radiation Long After Exposure
Biosciences researchers have established a powerful new method that couples advanced infrared imaging techniques with statistical machine learning models to quickly and non-invasively determine with high accuracy whether an animal was exposed to radiation—even at extremely low doses almost three months post exposure.
Biography
Education & Training
PhD, Duke University, 1982
Research Interests
We focus on structural biology methods and applications for analysis and design projects that concern fundamental questions of molecular cell biology and biochemistry relevant to biological mechanisms and human disease.
DNA damage responses provide master keys to unlock oncogenic synthetic lethality and activate innate immunity for cancer biology and medicine going forward. Our research focuses on applying structural biochemistry and biophysics to molecular and cellular oncology and cancer biology for predictive molecular mechanisms and innovative strategies that target DNA damage responses in cancer. By solving and building upon >350 macromolecular structures, our group is developing and applying methods for integrating X-ray and cryo-EM data to build quantitative and foundational knowledge resulting in publications and patents aimed at advanced patient care.
Programs & Initiatives
Recent Publications
Related News
Time-Resolved SAXS Screen of Small-molecule Drug Candidates
A team of researchers developed a high-throughput drug-discovery workflow leveraging time-resolved small-angle X-ray scattering (SAXS) capabilities at the Advanced Light Source’s (ALS) Structurally Integrated Biology for the Life Sciences (SIBYLS) beamline to identify small molecules capable of activating biomolecular dynamics associated with a desired therapeutic outcome.
Enigmatic Protein Sculpts DNA to Repair Damage
Biosciences Area researchers and their collaborators have determined how a protein called XPG binds to and reshapes damaged DNA, illuminating its role in averting genetic disease and cancer.
Programming Proteins to Pair Perfectly
Bioscientists at the Advanced Light Source (ALS) at Berkeley Lab lent their expertise to a project led by scientists at the University of Washington to design proteins in the lab that zip together like DNA. The technique could enable the design of protein nanomachines to help diagnose and treat disease, allow for more precise engineering of cells, and perform a variety of other tasks.
Research Interests
My lab is interested in the general question of how signaling circuits generate complex cellular behaviors such as cell polarity, motility, growth, and differentiation. Because these processes have been difficult to understand with standard techniques, we are developing new approaches to isolate and quantitatively interrogate individual steps in the signaling cascade. We are particularly interested in four areas:
1. Understanding the self-organizing circuits that underlie cell polarity, directed migration, and morphogenesis.
2. Developing optogenetic tools to probe the spatial and temporal logic of signal transduction cascades.
3. Investigating how mechanotransduction integrates the local and global organization of migrating cells.
4. Probing how the three-dimensional structure of the eukaryotic genome regulates gene expression.
Recent Publications
Divisions
- Science Programs
Secondary Affiliation:
Environmental Genomics and Systems Biology
- Comparative and Functional Genomics
Recent Publications
Related News
University of Duisburg-Essen Delegation Explores Collaborative Opportunities with Lawrence Berkeley National Laboratory
Building on a Memorandum of Understanding signed between UDE and Berkeley Lab researchers, a kick-off meeting focused on future collaborations in the fields of genomics, structural biology, bioimaging, and water research.
JGI-Enabled Research Finds Flagella in the Terrestrial Roots of Marine Bacteria
Ancient Chloroflexotas ditched their flagella and other traits when migrating back to the ocean.
Studying the Tiniest Archaea, JGI Users Find a Genomic Switch of Friend or Foe
Meta-omics datasets show that CRISPR-Cas systems determine mutualism or parasitism between some archaeal hosts and their hitchhikers.
Recent Publications
Related News
Clues to How Space Radiation Induces Cognitive Damage
The documented mechanism for radiation-induced health risks like cancer has typically been damage to the DNA contained in the nucleus of cells. But a recent analysis suggests that radiation by certain light ions, which abound in outer space, instead disrupt a separate cellular structure, smaller than a cell nucleus or a synapse.
To Find Mutated Sperm, Go FISH
Chemotherapy and radiation treatments can be life-saving for patients with cancer, but they have harsh side effects that can been felt and seen throughout the body. There can also be unseen consequences: These important treatments can mutate DNA and damage chromosomes in patients’ cancerous and noncancerous cells alike. When this occurs in a germline cell (eggs in women and sperm in men), it can lead to serious fetal and birth defects in a resulting pregnancy. In a study published in PLOS One, a team led by Biological Systems and Engineering (BSE) Division senior scientist Andrew Wyrobek reported success adapting an established cellular DNA analysis technique called fluorescence in situ hybridization (FISH) to probe sperm DNA for a wide variety of chromosomal defects simultaneously.
BRAIN Neuro-Workshop Report Now Available
The report on the multi-institutional Neuro-Workshop held June 2, 2016 at Berkeley Lab (LBNL) is now available. The workshop was convened to discuss relevant technological capabilities for the national Brain Research through Advancing Innovative Neurotechnologies (BRAIN) Initiative. Participants also explored collaborative opportunities that would address neuroscience “grand challenges” in support of the Department of Energy (DOE) contribution to BRAIN and other national scientific challenges.