Matthew Blow
Data Science Supervisor
Divisions
- Genomic Technologies
Secondary Affiliation:
Environmental Genomics and Systems Biology
- Comparative and Functional Genomics
Research Interests
Using sequence-based approaches to understand genome function.
Recent Publications
Related News
Biosciences Area FY16 LDRD Projects
The projects of eleven Biosciences Area scientists and engineers received funding through the FY2016 Laboratory Directed Research and Development (LDRD) program. These projects cover a broad range of topics, including energy science technology applications, novel computing technologies, and mechanistic understanding of multi-scale interactions among molecules, microbes, plants, metazoans, the abiotic environment, and their feedbacks. Together, these efforts account for nearly 14% of the $25.3 million allocated. Lab-wide, 84 proposals were selected from a field of 179.

Building: 33, Room 349
Mail Stop: 33R0345
Phone: (510) 486-5709
Fax: (510) 486-5909
BKPoon@lbl.gov
https://phenix-online.org
https://cci.lbl.gov
Links
Programs & Initiatives
Recent Publications
Related News
Cracking the Code: Using AI to Solve Difficult-to-map Proteins
Leveraging artificial intelligence and quantum calculations, scientists developed a new tool that yielded higher-quality structural information and solved notoriously elusive proteins.
Congratulations to Biosciences Area Director’s Award Recipients
Numerous Biosciences Area personnel are among the 2021 Berkeley Lab Director’s Awards honorees. This annual program recognizes outstanding contributions by employees to all facets of Lab activities. A complete list of winners can be found here. The 10th annual Director’s Awards ceremony will take place on November 18 at noon.

Building: Donner Laboratory, Room 351A
Mail Stop: DONNER
Phone: (510) 708-2564
SHKim@lbl.gov
https://chemistry.berkeley.edu/faculty/chem/kim
Links
Divisions
Molecular Biophysics and Integrated Bioimaging
- Structural Biology
Secondary Affiliation:
Environmental Genomics and Systems Biology
- Biosystems Data Science
Research Interests
A. Construction of whole genome phylogeny of living organisms, “Tree of Life”
The first task for this project is to develop one or more methods for comparing whole genome sequences of two organisms, not just a set of highly conserved gene or protein sequences, as currently practiced in Multiple Sequence Alignment (MSA) method. Our starting point was treating each whole genome sequence as a book consisting of a single string of alphabets without spaces between words for each chromosome. My group has developed the “Feature Frequency Profile (FFP)” method, which is a variation of “Word Frequency Profile” method used to compare two books describe in the field of Natural Language Analysis. Using the FFP method we were able to construct phylogenic trees of three domains of Life at an intermediate resolution, and two of the most diverse and large groups of Life, Prokaryotes (Archaea and Bacteria combined), and Fungi, the largest kingdom of Eukarya, at a high resolution. Compared to those trees based on MSA methods, our results revealed high similarities in grouping (clading) at high phylogenic levels, but substantial differences in evolutionary branching order of the clades at deeper evolutionary levels. Our next projects are to construct the phylogenic trees of other large “phylogenic” groups such as protists, Eukaryotic algae, insects, plants and others, and ultimately “Tree of Life” for all living organisms for which whole genome sequences are available.
B. Whole genome variation of human species vs. disease susceptibility
Most regions of genomes of normal human cells have been found to have the same sequences among individuals, but a small fraction, spread throughout the genome, have variations within a population. Of these, the single nucleotide polymorphisms (SNPs) account for the largest number of variations and, have been identified in over 3 million genomic “tag” positions out of 3 billion positions in a whole haploid genome. It has been widely accepted that the analysis of SNPs may be able to allow one to predict the genomic component of the susceptibility of individuals to complex diseases such as cancers, neurological diseases, autoimmune diseases, and other traits. So far, the results from the current analysis methods (e.g. Genome-wide Association Studies method) and interpretation of them have yielded information of limited predictive value of practical utility for making health-related decisions at individual or population level without information of family histories.
Recognizing the complexity and heterogeneity of cancer mechanisms, we have developed, using SNPs, an empirical approach using supervised machine-learning method, a branch of Artificial Intelligence, for predicting the relative genomic susceptibility of an individual to 9 traits consisting of 8 major cancer classes plus a healthy class. The multiclass accuracy of the combined prediction ranges from 33 to 56% depending on cancer classes of testing sets, as compared to 11% for a random prediction among 9 traits. Despite limited SNP data available and absence of rare SNPs in public databases at present, the results suggest that the framework of this approach or its improvement can predict the cancer susceptibility with probability estimates useful for making health-decisions for individuals or for a population. Our next projects are to use similar approaches to predict genomic susceptibility for various neurological diseases and autoimmune diseases.
C. Whole genome variation of non-human species vs. traits
For a longer-term projects we plan to apply similar approaches of machine learning methods as in B above to the genomic variations of various non-human species such as crop and bio-fuel plants, insects, farm animals and other to predict traits such as drought resistance, high growth, insect resistance, disease resistance etc.
Biography
Dr. Kim is a Professor of Graduate Studies in the Department of Chemistry, and a faculty member of Center for Computational Biology, University of California, Berkeley, CA, USA. He is also a Faculty Affiliate, Division of Molecular Biophysics and Integrated Bioimaging, Lawrence Berkeley National Laboratory, Berkeley, CA. His research area has been in Structural Biology of signal communicating proteins, such as Ras, and protein kinases that signal many cellular processes, such as cell differentiation and cancer. More recently his group have been applying machine learning methods for predicting individual’s genomic susceptibility for complex diseases such as cancer using whole genome germline sequence variations. More recently, his group has developed a new computation method to construct a “Whole-genome Tree of Life” that reveals kinship among all living organisms using whole-genome sequence information based on Information Theory. A similar approach is being used to study genomic demography of human ethnic groups. He is a member of the US National Academy of Sciences, a Fellow of the American Academy of Arts and Sciences, and a Fellow of The American Association for the Advancement of Science.
Recent Publications
Related News
Two Bioscientists Among Those Named AAAS Fellows
Two scientists from the Biosciences Area, Sung-Hou Kim and Susannah Tringe, have been named Fellows of the American Association for the Advancement of Science (AAAS). They join fellow Lab scientists Allen Goldsten, faculty scientist in the Energy Technologies Area, and Kathy Yelick, associate laboratory director of Computing Sciences, in receiving the distinction of Fellow this year for “their scientifically or socially distinguished efforts to advance science or its applications.”
Research Interests
Chemical accuracy in macromolecular Xray crystallography
Recent Publications
Related News
Cracking the Code: Using AI to Solve Difficult-to-map Proteins
Leveraging artificial intelligence and quantum calculations, scientists developed a new tool that yielded higher-quality structural information and solved notoriously elusive proteins.
Congratulations to Biosciences Area Director’s Award Recipients
Several Biosciences Area personnel are among the 2024 recipients of Berkeley Lab Director’s Achievement Awards. The program recognizes outstanding contributions by employees to all aspects of Lab activities.
Nigel Moriarty, Wave Wizard
Experimentation abounds for computational research scientist Nigel Moriarty. A lifelong stamp collector and surfer, he approaches culturally-distinct pockets of society with the curiosity of an anthropologist. And applying quantum chemistry and theoretical physics to drug design is part of his day-to-day work writing software with MBIB’s PHENIX group.

Building: 70A, Room 3317L
Mail Stop: 70A-3317
Phone: (510) 486-5943
Fax: (510) 486-7152
HYHolman@lbl.gov
https://bsisb.lbl.gov
Links
Biography
Hoi-Ying Holman is a senior scientist in the Molecular Biophysics and Integrated Bioimaging (MBIB) Division of Berkeley Lab’s Biosciences Area and Director of the Berkeley Synchrotron Infrared Structural Biology (BSISB) imaging program at the Advanced Light Source (ALS). She received her PhD from the University of California at Berkeley. With team members Drs. Sun Choi, Liang Chen, and Giovanni Birarda, she was awarded a 2014 R&D 100 award for their development of the Berkeley Lab Multiplex Chemotyping Microarray (MCM).
Research Interests
Hoi-Ying’s research interests are in microbial ecology, geomicrobiology, bioenergy, bioremediation, and bioavailability of chemicals to ecological receptors. In the BSISB imaging program at the ALS, she focuses on developing and providing research communities new synchrotron infrared (SIR) technologies for deciphering the relationship between genome and functional processes, and identifying the connection between the genome and natural environments. Current technologies in active development are: SIR nano-spectroscopy, SIR plasmon resonance microscopy, and the integration of SIR spectromicroscopy with ambient IR laser ablation (AIRLAB) mass spectrometry.
Programs & Initiatives
Recent Publications
Related News
University of Duisburg-Essen Delegation Explores Collaborative Opportunities with Lawrence Berkeley National Laboratory
Building on a Memorandum of Understanding signed between UDE and Berkeley Lab researchers, a kick-off meeting focused on future collaborations in the fields of genomics, structural biology, bioimaging, and water research.
Watching the Enzymes that Convert Plant Fiber into Simple Sugars
Researchers in the Berkeley Synchrotron Infrared Structural Biology (BSISB) Imaging Program developed a technique that combines a novel microfluidic device and infrared spectroscopy to study how a cellulose-degrading enzyme works in real time.
Whip It: Novel Liquid Jet Makes Droplets March to the Beat
An interdisciplinary team has developed a first-of-its-kind, steady-state whipping liquid microjet that produces droplets of uniform size and spacing in a two-dimensional profile. The technology could ultimately lead to advancements in structural biology, climate science, and several industries.

Building: 33, Room 243
Mail Stop: 33R0229
Phone: (510) 486-4988
Fax: (510) 486-6880
SETsutakawa@lbl.gov
Recent Publications
Related News
Biosciences FY26 LDRD Projects
The Laboratory Directed Research and Development program at Berkeley Lab produces cutting-edge research for the DOE and the nation. Read about the Biosciences Area–led projects and multi-Area collaborations with Biosciences co-investigators receiving funding this cycle.
New Statistical Technique Improves Predictions of Protein Flexibility
Computational and AI methods for structural biology prediction fall short in the face of proteins that morph as part of their function. A team of researchers at Duke University and Berkeley Lab defined the problem and devised statistical techniques to improve the models’ accuracy.
Biosciences FY25 LDRD Projects
The projects of 23 Biosciences Area scientists and engineers received funding through the FY25 Laboratory Directed Research and Development (LDRD) program.

Building: 978, Room 4466
Mail Stop: 978-4121
JCMortimer@lbl.gov
http://www.mortimerlab.org/home.html
https://www.jbei.org/person/jenny-mortimer/
https://www.sorghummetabolicatlas.org/
https://researchers.adelaide.edu.au/profile/jenny.mortimer
Links
Divisions
Environmental Genomics and Systems Biology
- Molecular EcoSystems Biology
Secondary Affiliation:
Biological Systems and Engineering
- BioEngineering & BioMedical Sciences
Research Interests
Jenny is a plant glycobiologist who is interested in understanding the myriad ways that plants synthesize and use complex sugars: to build their cell wall, and to glycosylate other molecules, including proteins and lipids. Her team applys synthetic and systems biology tools with the overal goal to develop more sustainable bioenergy crops.
As part of the Joint BioEnergy Institute (JBEI) we are deciphering how a cell wall is made and assembled, and applying this knowledge to the predictable engineering of dedicated biomass crops (primarily sorghum and switchgrass). As part of the m-CAFEs SFA (Microbial Community Analysis & Functional Evaluation in Soils Scientific Focus Area), we are exploring how roots interact with a synthetic microbial community. We aim to enhance those interactions via engineering, to support the development of more sustainable crops that require fewer inputs (e.g. irrigation, pesticides). For m-CAFEs, and other projects, we are making use of fabricated ecosystems (EcoFABs, EcoPODS), as we attempt to bridge the gap between data gathered in the lab and that collected in the field. Other projects include exploring how cell wall esters contribute to forest volatile emissions when trees undergo drought, and developing resources to support sorghum’s use in biotechnology research.
Programs & Initiatives
Recent Publications
Related News
Plant Single-cell Solutions for Energy and the Environment Workshop Report Released
On January 23, 2020, Berkeley Lab hosted a workshop on opportunities afforded by single-cell technologies for energy and environmental science, as well as conceptual and technological grand challenges that must be tackled to apply these powerful approaches to plants, fungi and algae. This event, which was spearheaded by Diane Dickel in the Environmental Genomics and Systems Biology Division, brought together a diverse group of leaders in functional genomics technologies from academia, the National Laboratories, and local research institutions.
Biosciences Area FY21 LDRD Projects
The projects of 15 Biosciences Area scientists and engineers received funding through the FY21 Laboratory Directed Research and Development (LDRD) program.
Mortimer Participates at AAAS Forum on Science & Technology Policy
 Jenny Mortimer, Deputy Vice President of the Feedstocks Division at the Joint BioEnergy Institute (JBEI) and Scientist with the Environmental Genomics and Systems Biology (EGSB) Division, participated at a 2018 AAAS Forum on Science & Technology Policy panel entitled “Science Competitiveness in Relation to Public Support for Science”. Panelists discussed how the scientific community must work to maintain societal relevance and build trust. Mortimer presented a code of ethics for scientists recently developed by the World Economic Forum’s Young Scientists community. The code serves as a tool to nurture a positive change of culture in the research world by not only guiding and shaping the behavior of individuals but also the processes of the scientific institutions that are to facilitate this cultural shift.
Jenny Mortimer, Deputy Vice President of the Feedstocks Division at the Joint BioEnergy Institute (JBEI) and Scientist with the Environmental Genomics and Systems Biology (EGSB) Division, participated at a 2018 AAAS Forum on Science & Technology Policy panel entitled “Science Competitiveness in Relation to Public Support for Science”. Panelists discussed how the scientific community must work to maintain societal relevance and build trust. Mortimer presented a code of ethics for scientists recently developed by the World Economic Forum’s Young Scientists community. The code serves as a tool to nurture a positive change of culture in the research world by not only guiding and shaping the behavior of individuals but also the processes of the scientific institutions that are to facilitate this cultural shift.

Building: 2151 Berkeley Way, Room 512G
Phone: (510) 643-0113
JDoudna@lbl.gov
http://rna.berkeley.edu
Links
Research Interests
Molecular and Cell Biology and Chemistry
Recent Publications
Related News
Doudna Elected to the National Academy of Engineering
MBIB faculty scientist Jennifer Doudna was named to the Academy's 2026 class of new members in recognition of her highly impactful work developing gene editing methods based on CRISPR-Cas9.
Doudna Awarded American Chemical Society’s Priestley Medal
Jennifer Doudna is the 2026 recipient of the American Chemical Society's Priestley Medal. She is honored for outstanding discoveries on ribozyme function, Dicer and double-stranded RNA processing and CRISPR gene editing, and for impactful international science leadership.
How Berkeley Lab is Leading the Biology-Based Industrial Revolution
Biosciences Area researchers are playing a key role in shaping the future of biomanufacturing.

Building: 55, Room 219
Mail Stop: 55R0121
Phone: (510) 486-5065
Fax: (510) 486-4768
WJJagust@lbl.gov
http://jagustlab.neuro.berkeley.edu
Recent Publications
Related News
Regional Tau Deposits Predict Alzheimer Disease
Berkeley Lab researchers propose a new model for the early pathology of this debilitating Alzheimer disease.
UCB Study Finds Sleep May Be a Biomarker for Dementia
Research led by UC Berkeley scientists found that adults who reported a decline in sleep quality in midlife (40s–60s) had more beta amyloid and tau clusters in their brains—both of which are associated with a higher risk of developing dementia later in life. The same study also revealed that people with high levels of tau protein in their brains were more likely to lack the synchronized brain waves that are crucial to getting a good night’s sleep. Together, the findings suggest that sleep changes detectable in a simple overnight sleep study may serve as biomarkers for later risk of dementia.
Jagust Wins Radical Ideas in Brain Science Challenge
Congratulations to William Jagust, senior faculty scientist in the Molecular Biophysics and Integrated Bioimaging Division, for winning the 2018 Radical Ideas in Brain Science Challenge, made possible through the generosity of Berkeley Brain Initiative donors. Jagust, who is also Professor of Public Health at UC Berkeley, will receive up to $190,000 over two years to investigate the degradation of the blood-brain barrier as a potential paradigm-shifting culprit in Alzheimer’s disease and other dementias.
Research Interests
Dr. Chris Mungall is a Staff Scientist in the Environmental Genomics and Systems Biology Division at LBNL, where he heads the Biosystems Data Science department. Chris’s research interests center around the capture, computational integration, and dissemination of biological research data, and the development of methods for using this data to elucidate biological mechanisms underpinning the health of humans and of the planet. He and his team have led the creation of key biological ontologies for the integration of resources covering gene function, anatomy, phenotypes and the environment. In the Gene Ontology project and others, Chris and his collaborators develop systems that help curators translate biological knowledge into a computable form, and apply that biological knowledge to answer complex biological questions.
Chris’s areas of focus include ontologies, systems biology, data science, biocuration, knowledge representation, data harmonization, reusable and interoperable software, machine learning and reasoning. A growing area of interest is translating curated basic research data into clinically actionable frameworks.
Chris is a PI on the Gene Ontology (GO), the Monarch Initiative, the Alliance of Genome Resources, Phenomics First, and the NCATS Biomedical Data Translator, as well as metadata lead for the National Microbiome Data Collaborative (NMDC). In 2017, Chris was the first person to be awarded the Exceptional Contributions to Biocuration Award by the International Society for Biocuration. In 2020, he received a Berkeley Lab Early Scientific Career Director’s Award.
Programs & Initiatives
Recent Publications
Related News
Biosciences FY25 LDRD Projects
The projects of 23 Biosciences Area scientists and engineers received funding through the FY25 Laboratory Directed Research and Development (LDRD) program.
Researchers Aid in Quest to Identify GMOs
Biosciences Area researchers led testing and evaluation of technologies developed to quickly distinguish genetically modified organisms from naturally occurring ones. They designed and produced biological samples of increasing complexity to assess how well the tools performed.
EGSB Researchers Tapped for Bridge2AI
A team headed by the Environmental Genomics and Systems Biology (EGSB) Division's Chris Mungall, will be part of the National Institute of Health program, Bridge to Artificial Intelligence (Bridge2AI). Mungall and his colleagues will be collaborators in the Standards Core, led by the University of Colorado.

Building: 6, Room 2104
Mail Stop: 6R2100
Phone: 510 457 6317
MHammel@lbl.gov
https://bl1231.als.lbl.gov/
Links
Research Interests
My early scientific career began with the characterization of drug delivery in Low-Density Lipoprotein and mastering solution X-ray scattering (SAXS) techniques at the Austrian Academy of Sciences in Graz, Austria. I have been bridging crystallography with solution scattering since 2000. My early developments in integrative structural biology were later awarded Erwin Schrödinger fellowship to pursue my postdoctoral research in Marseille at Centre National de la Recherche Scientifique. Here I developed an ensemble modeling approach to investigate flexible and dynamic macromolecules by combining solution scattering, crystallography, and molecular dynamics. This early innovation becomes the groundwork for our current worldwide used FoXS software package.
Besides developing novel SAXS methods, my research focuses on visualizing the large megadalton biological assemblies that included the cellulosome, human complement, and non-homologous end-joining. Characterizing these dynamic molecules impacts medical research, like defining the allosteric mechanism for complement inhibition by the Staphylococcus aureus and discovering the extended grooved scaffold for DNA ligation in DNA break repair.
My effort as the beamline scientist at the SIBYLS beamline at the Advanced Light Source (ALS) makes the SAXS technique accessible to more investigators and pushes SAXS onto a new paradigm. We have been the first to create a high throughput SAXS pipeline and established a mail-in program at SIBYLS to collect SAXS data for scientists from all around the world.
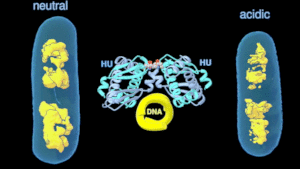
As a research scientist at the LBNL, my research focuses on the modulation of transcription by DNA control elements. Using an altered bacterial histone-like protein, we show that reorganization of the bacterial chromatin can dynamically modulate the cellular transcription pattern. We anticipate making histone-like protein interactions an attractive target for controlling pathogenesis and microbial systems in general with this ongoing research.
Change in the architecture of the bacterial chromosome during the adaptation to an acidic environment is controlled by the DNA binding protein called HU and its interaction with DNA. (Credit: Michal Hammel/Berkeley Lab)
I’m co-directing Structural Cell Biology Core of the NCI-funded Structural Cell Biology of DNA Repair (SBDR) group.
Recent Publications
Related News
SIBYLS Team Recognized with Innovative Instrumentation Award
The Structurally Integrated BiologY for the Life Sciences (SIBYLS) team received the 2025 Klaus Halbach Award for Innovative Instrumentation at the 2025 Advanced Light Source (ALS) User Meeting in August.
A Fast Track for Visualizing RNA Structures
Scientists have combined multiple AI tools into a single streamlined process that predicts the atomic structures of RNA molecules.
Researchers Leverage SAXS to Understand Aspect of Microbial Metabolism
A team of Molecular Biophysics and Integrated Bioimaging Division researchers used synchrotron technology unique to the Advanced Light Source (ALS) at Berkeley Lab to probe the conformational states behind electron bifurcation.

Building: 66, Room 308
Mail Stop: 66
Phone: (510) 486-4325
Fax: (510) 486-4995
HMFrei@lbl.gov
http://go.lbl.gov/teachingmodules
Links
Research Interests
The goal of our work is to develop efficient, robust photocatalytic subsystems and complete systems for the synthesis of renewable fuels and chemicals using carbon dioxide and water as starting materials, and sunlight as energy source. All-inorganic oxo-bridged heterobinuclear light absorbers coupled to metal oxide nanoclusters are being developed for accomplishing visible light induced multi-electron catalysis for carbon dioxide reduction and water oxidation, using photodeposition methods for the proper coupling of chromophore and nanocluster catalyst. Structures, charge transfer processes and catalytic mechanisms are elucidated by FT-infrared, Raman and optical spectroscopy, X-ray spectroscopy, and atomic resolution imaging. The mechanistic understanding gained from the time-resolved FT-infrared studies combined with electron transfer investigations of the heterobinuclear charge transfer units by transient optical spectroscopy guide the design of units for improved photocatalytic efficiency under visible and near infrared light. Using low temperature atomic layer deposition and nanofabrication methods, metal oxide core-shell constructs are being developed for separating the water oxidation catalysis from light absorption and reduction chemistry by a nanoscale silica-based membrane. Synthetic methods have been established for embedding electron or hole conducting molecular wires into the insulating silica membrane, and proton transmission properties of the silica have been quantified. Emphasis is on tightly controlled electron transport from light absorber to catalyst through the molecular wires, on atomically defined contacts between the components, and on the elucidation of charge transport kinetics and efficiency across the assembly. The long term objective is to close the photocatalytic cycle of carbon dioxide reduction and water oxidation on the nanoscale while achieving product separation on the macroscale.
The ultrathin silica membranes with embedded molecular wires open up opportunities for exploring single integrated biohybrid assemblies to create function that combines the best of biology with the best of inorganic catalysis (BSP milestone). The membranes allow electronic coupling of life organisms with inorganic catalysis, with the incompatible reaction environments held in nanometer proximity. The approach drastically reduces efficiency losses due to transport of charges and chemical species over macroscale distances. Major effort will focus on spectroscopic and electrochemical studies to understand and control electron and proton transport pathways across both the silica and biological membrane, and to elucidate the catalytic mechanisms.
Recent Publications
Related News
Nature-Inspired Green Energy Technology Clears Major Development Hurdle
Heinz Frei, a senior scientist in Biosciences’ Molecular Biophysics and Integrated Bioimaging (MBIB) Division, seeks to engineer devices that emulate photosynthesis – the sunlight-driven chemical reaction that green plants and algae use to convert carbon dioxide (CO2) into cellular fuel. If the necessary technology could be refined past theoretical models and lab-scale prototypes, this idea, known as artificial photosynthesis, has the potential to generate large sources of completely renewable energy using the surplus CO2 in our atmosphere.
Ultrathin Membrane Both Isolates and Couples Living and Non-Living Catalysts
Biosciences researchers have developed a novel nanoscale membrane embedded with molecular wires that simultaneously chemically isolates, yet electrochemically couples, a microbial and an inorganic catalyst on the shortest possible length scale. This new modular architecture, described in a paper recently published in Nature Communications, opens up a large design space for building scalable biohybrid electrochemical systems for a variety of applications.
A Core−Shell Nanotube Array for Artificial Photosynthesis
“The key design principle of natural photosynthesis is the closing of the photosynthetic cycle on the shortest possible length scale under membrane separation of the incompatible water oxidation and proton reduction environments,” said Heinz Frei, a senior scientist in Biosciences’ Molecular Biophysics and Integrated Bioimaging (MBIB) Division. With collaborators Eran Edri, a former postdoctoral fellow in MBIB now at Ben-Gurion University, and Shaul Aloni in the Molecular Foundry Division, Frei developed a fabrication method to make a square-inch sized artificial photosystem, in the form of an inorganic core-shell nanotube array, that implements this design principle for the first time. The method was described in a paper published earlier this year in ACS Nano.