Amy Herr is one of three recipients of the Berkeley Chamber of Commerce’s Visionary of the Year award for 2017. The honor is bestowed annually to local innovators tackling real-world challenges with “imagination and persistence.” Herr is a professor of bioengineering at UC Berkeley and a Berkeley Lab Biosciences Area faculty engineer with a primary appointment in Biological Systems and Engineering (BSE) and a secondary appointment in Molecular Biophysics and Integrated Bioimaging (MBIB). Her research focus is on inventing tools to analyze the levels of various proteins within single cells, which has applications for the treatment of diseases such as cancer. Read more from UC Berkeley News.
American Society for Microbiology Honors Nikos Kyrpides
Nikos Kyrpides, head of the Prokaryote Super Program at the Department of Energy Joint Genome Institute (DOE JGI) and a senior scientist in the Biosciences Environmental Genomics and Systems Biology (EGSB) division, was selected as the 2018 USFCC/J. Roger Porter Award recipient by the American Society for Microbiology (ASM). He will be honored at the 2018 ASM Microbe Meeting in Atlanta, Ga. The award recognizes outstanding effort by a scientist in demonstrating the importance of microbial biodiversity through sustained curation or stewardship of a major resource used by the scientific community. For more than a decade, Kyrpides and his colleagues have been working toward the goal of developing a comprehensive catalog of reference genomes for every bacterial and archaeal species. Read more from JGI.
X-ray Footprinting Reveals Secrets of ‘Metal-Breathing’ Bacterium
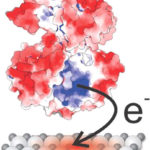
A team of Berkeley Lab researchers conducted X-ray footprinting mass spectrometry (XFMS) experiments at the Lab’s Advanced Light Source (ALS) to pinpoint how a protein of the bacterium Shewanella oneidensis transfers electrons to a metal oxide substrate. The research was led by Caroline Ajo-Franklin, whose lab is part of the Molecular Foundry and who holds a secondary appointment in the Molecular Biophysics and Integrated Bioimaging (MBIB) division, in collaboration with Corie Ralston, also of MBIB. Tatsuya Fukushima, a former postdoc in Ajo-Franklin’s lab, and Sayan Gupta, a member of Ralston’s lab, were co-first authors on the paper published in the Journal of the American Chemical Society. The study, which identified an unexpectedly small and weak binding site, also benefitted from expertise and tools contributed by Joint BioEnergy Institute (JBEI) and Biological Systems and Engineering (BSE) researchers Christopher Petzold and Leanne Jade Chan. Read more at the Berkeley Lab News Center.
Bay Area Biopharma Company Uses ALS to Tackle Sickle Cell Disease
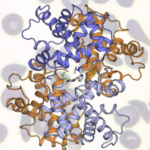
The protein crystallography capabilities at the Advanced Light Source’s (ALS’s) Beamline 8.3.1 have been critical to Global Blood Therapeutics’ (GBT’s) ongoing effort to formulate a better treatment for sickle cell disease (SCD).
Congressman Randy Hultgren Visits Biosciences Area
 U.S. Congressman Randy Hultgren (R-IL-14) visited the Biosciences Area’s Emery Station Operations Center on September 1st. Congressman Hultgren met with Mary Maxon, Biosciences Associate Lab Director, and Jay Keasling, JBEI’s Chief Executive Officer, and also toured JBEI’s laboratories.
U.S. Congressman Randy Hultgren (R-IL-14) visited the Biosciences Area’s Emery Station Operations Center on September 1st. Congressman Hultgren met with Mary Maxon, Biosciences Associate Lab Director, and Jay Keasling, JBEI’s Chief Executive Officer, and also toured JBEI’s laboratories.
Hultgren is co-founder and leading member of the Science and National Labs Caucus and a proponent of science’s role in unlocking the potential for economic growth and job creation. He recently spoke at a Council on Competitiveness-sponsored event in Washington D.C. during which the Council released its latest report “Leverage: Advancing U.S. Bioscience”. The report focused on leveraging U.S. potential in the growing sector of bioscience and biomanufacturing.
- « Previous Page
- 1
- …
- 142
- 143
- 144
- 145
- 146
- …
- 213
- Next Page »
Was this page useful?