Antioch High School juniors who are taking part in the Biotech Partners program, are getting ready for their interviews in order to get summer internships. Emery Station Operations Center’s operational staff was on hand this week to help the students practice their job application skills during mock interviews.
Antioch High School juniors who are taking part in the Biotech Partners program, are getting ready for their interviews in order to get summer internships. Emery Station Operations Center’s operational staff was on hand this week to help the students practice their job application skills during mock interviews.
New Insight about Antidepressants
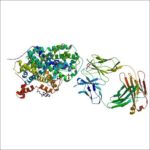
Researchers at the Oregon Health & Science University (OHSU) Vollum Institute have discovered how chemically diverse drugs used to treat depression and anxiety disorders interact with the protein that transports serotonin in the brain. Jonathan Coleman and Eric Gouaux used X-ray crystallography techniques at two U.S. Department of Energy national user facilities to collect data: Molecular research was carried out at the Biosciences’ Berkeley Center for Structural Biology, beamline 5.0.2, located at the Advanced Light Source; and at the Advanced Photon Source (APS), at Argonne National Laboratory. Their findings, published in the January 29 issue of the journal Nature Structural & Molecular Biology, could open the way for the development of additional forms of antidepressants collectively known as selective serotonin reuptake inhibitors, or SSRIs.
Impact of Environmental Changes on Microbes in Arctic Soils
As the Arctic continues to warm at about twice the rate of the rest of the world, scientists expect its frozen soils—known as permafrost—to thaw, activating microbes capable of decomposing soil and releasing carbons and other nutrients to the atmosphere and water. Berkeley Lab scientists in the Earth and Environmental Systems Area (EESA), led by microbial ecologist Neslihan Taş, set out to learn more about how Arctic soil microbes can contribute to greenhouse gas emissions under a warming climate. Taş’s research team collaborated with Susannah Tringe, Deputy for User Programs at the Joint Genome Institute (JGI), to conduct their study, funded by the Department of Energy’s Office of Biological and Environmental Research (BER). The results were published in Nature Communications.
‘Minimalist’ Machine Learning Algorithms Analyze Images from Sparse Data
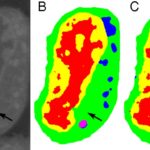
Typical machine learning methods used to analyze experimental imaging data rely on tens or hundreds of thousands of training images. But Daniël Pelt and James Sethian of Berkeley Lab’s Center for Advanced Mathematics for Energy Research Applications (CAMERA) have developed what they call a “Mixed-Scale Dense Convolution Neural Network” (MS-D) that “learns” much more quickly from a remarkably small training set. One promising application of MS-D is in understanding the internal structure and morphology of biological cells to identify, for example, differences between healthy and diseased cells. In one such project in Carolyn Larabell’s lab, the method needed data from just seven cells to determine the cell structure.
Using Nature’s Blueprint for Sustainable Indigo Dyeing Process
Indigo has been prized since antiquity for its vibrancy and deep blue hue and, for more than a century, its unique properties have been leveraged to produce the popular textile blue denim. However, the dyeing process requires chemical steps that are environmentally damaging. A team of researchers in the Molecular Biophysics and Integrated Bioimaging (MBIB) and Biological Systems and Engineering (BSE) Divisions, at JBEI, and UC Berkeley have developed a promising sustainable indigo dyeing process that relies on genetically engineered bacteria, mimicking the natural biochemical protecting group strategy employed by the Japanese indigo plant Polygonum tinctorium.
- « Previous Page
- 1
- …
- 130
- 131
- 132
- 133
- 134
- …
- 214
- Next Page »
Was this page useful?