Viruses and the organisms they infect are constantly working to outperform one another: the host evolves a way to defend and protect itself, the virus develops a way around and through those defenses, and so the cycle continues. For example, the viruses that infect soil bacteria (phages) can have a major effect on the plant microbiome. Recently, a collaborative effort involving researchers in the lab of Molecular Biophysics and Integrated Bioimaging (MBIB) Division faculty scientist and m-CAFEs investigator Jennifer Doudna revealed an especially creative way that viruses neutralize the immune response normally triggered by a host cell under invasion.
A large and diverse set of enzymes in a virus, known as 2H phosphodiesterases (2H PDEs), act like molecular scissors and cut the oligonucleotides in a host cell that are responsible for triggering an immune response. With the host cell’s immune response effectively silenced, the virus can invade and replicate.
“Viruses are constantly evolving and working to outsmart the defense systems of their hosts,” said Adam Deutschbauer, a senior scientist in the Environmental Genomics & Systems Biology (EGSB) Division. “And they affect many hosts including bacteria, humans, and plants, so identifying molecular mechanisms of immune evasion are crucial for our understanding of everything from nutrient cycling in the environment to human health. ”
This work on phosphodiesterase (PDE)-mediated oligonucleotide cleavage built on research previously led, in part, by Ben Adler and undergraduate researcher Kendall Hsieh and partially funded by the Microbial Community Analysis & Functional Evaluation (m-CAFEs) Scientific Focus Area. In 2025, these researchers investigated the activation genes behind a clever defense mechanism in bacterial cells that uses decoy defense molecules to trick the virus. This bacterial adaptation, dubbed Panoptes after the many-eyed watchman of Greek mythology, was used as a benchmark against which the more recently discovered viral defense strategy was compared to provide insights into the evolutionary dynamics of the arms race at the enzyme level.
“This really stemmed from intellectual curiosity,” said Adler, a postdoctoral researcher previously in the EGSB Division and currently a BRAINS Fellow with Speculative Technologies. “As we began peeling back layers of information, we realized that the previous discovery started to explain what otherwise mysterious phage genes were doing.
Our understanding of the battlefield between bacteria and their viruses is fundamentally redrawn, opening new opportunities for designing precision antimicrobial agents.”
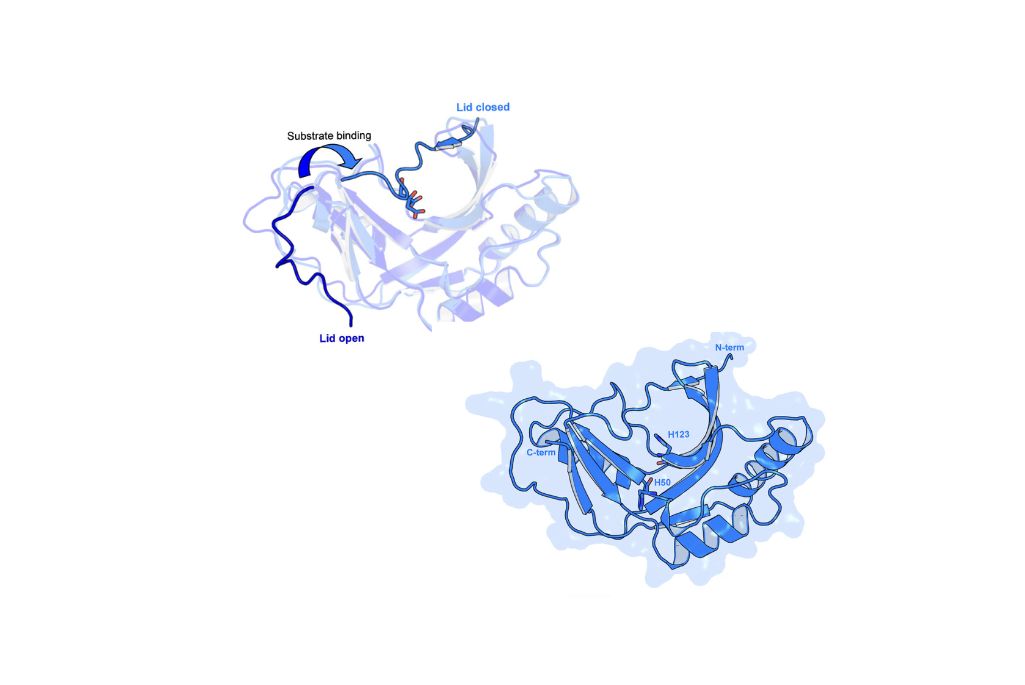
To understand the mechanics of the PDE-mediated viral silencing system, the researchers worked with beamline scientist Jay Nix at the Advanced Light Source (ALS). Using X-ray diffraction data collected at beamline 8.2.1 in the Berkeley Center for Structural Biology (BCSB), they solved the structures of multiple forms of 2H PDEs.
Doudna, senior author on the study, is also a professor at UC Berkeley, at the Innovative Genomics Institute (IGI), and is a Howard Hughes Medical Investigator.