Rising Sea Levels Could Mean Higher Wetlands Methane Emissions
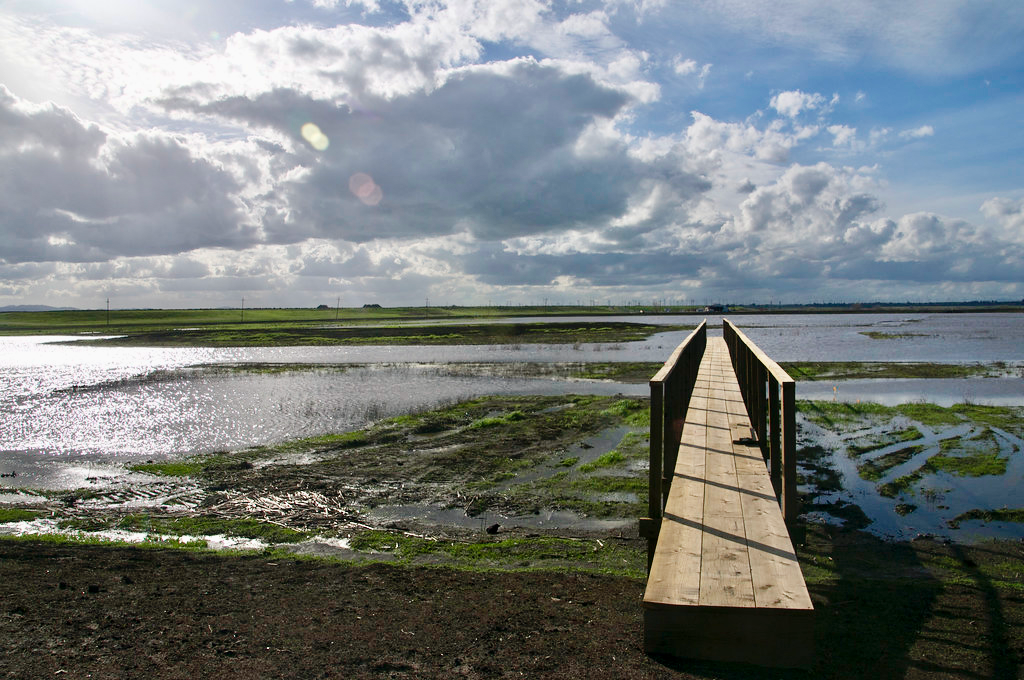

Area researchers led a team that examined the microbial, chemical, and geological features of 11 wetland zones in the Bay Area. Their findings indicate that the factors governing how much greenhouse gas is stored or emitted in natural landscapes are more complex and difficult to predict than previously thought.
More »