EcoFAB: A Tool for Combating Climate Change and Training the Next Generation
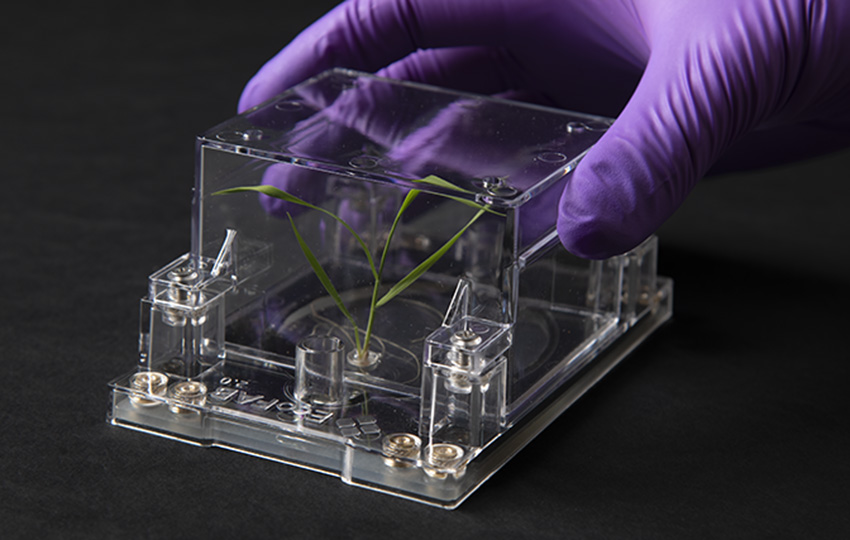

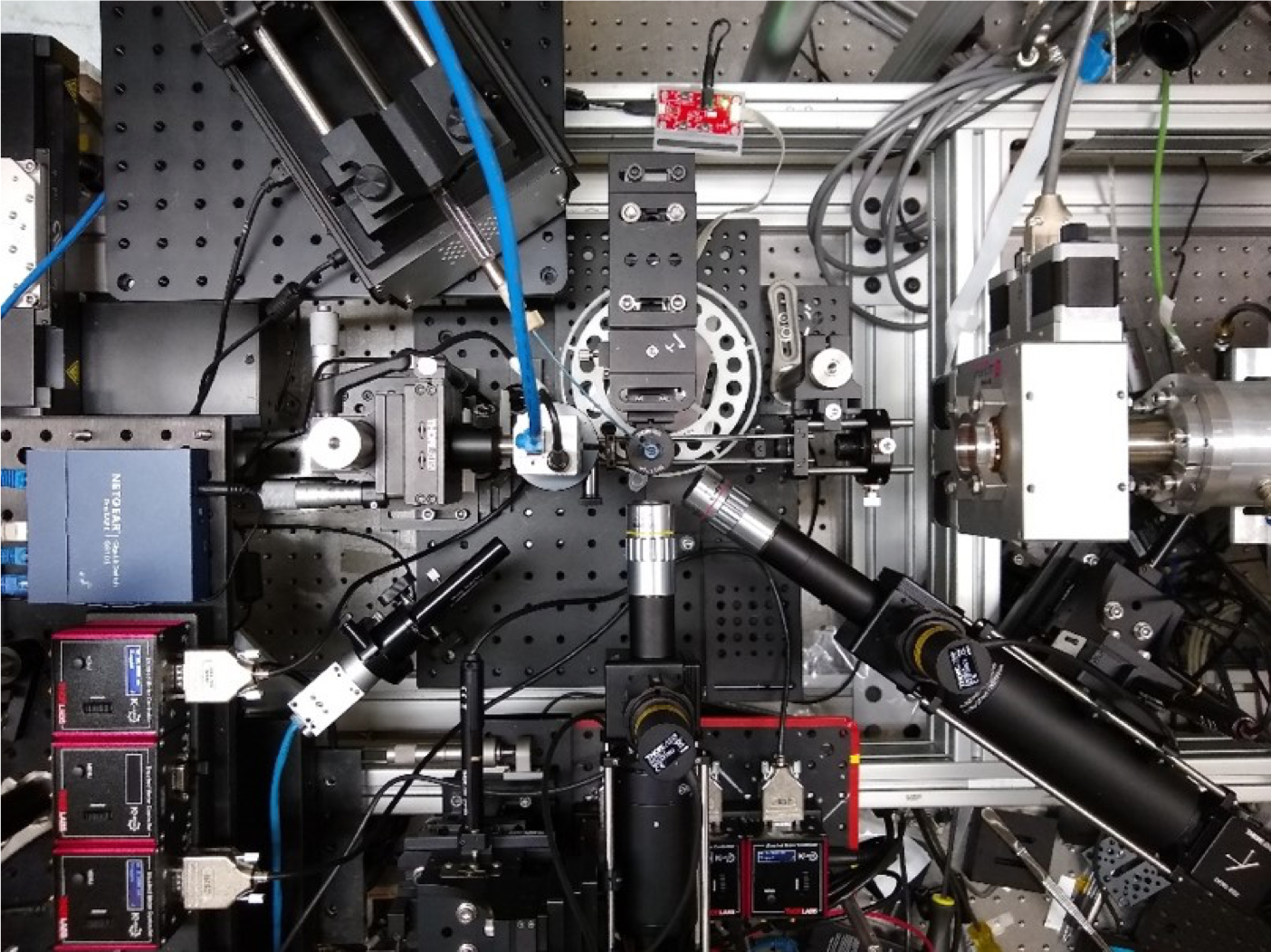
Fabricated ecosystems—EcoFABs—are plastic, takeout box–sized growth chambers developed at Berkeley Lab to be a standardized and reproducible platform for conducting experiments on model plants and the microbes that live around their roots. A greater understanding of how plants and microbes work together to store vast amounts of atmospheric carbon in the soil will help in the design of better bioenergy crops for the fight against climate change.
More »